 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Phvul.001G009800.1 | ||||||||
| Common Name | PHAVU_001G009800g | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Phaseolus
|
||||||||
| Family | SRS | ||||||||
| Protein Properties | Length: 334aa MW: 36016.5 Da PI: 7.7038 | ||||||||
| Description | SRS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | DUF702 | 223 | 6.1e-69 | 121 | 272 | 3 | 153 |
DUF702 3 sgtasCqdCGnqakkdCaheRCRtCCksrgfdCathvkstWvpaakrrerqqqlaaasskaaasa.aeaaskrkrelkskkq..salsstk 90
g+++CqdCGnqakkdC+h RCRtCCksrgf+C+thvkstWvpaakrrerqqqla+++++++ + + ++skr+re+ + ++ +l+++
Phvul.001G009800.1 121 GGGMNCQDCGNQAKKDCSHLRCRTCCKSRGFQCQTHVKSTWVPAAKRRERQQQLATLQQQNQPPQfRGDHSKRHRESIEGAAagGSLACAP 211
5789***************************************************998877655559999999999955544005566666 PP
DUF702 91 lssaeskkeletsslPeevsseavfrcvrvssvddgeeelaYqtavsigGhvfkGiLydqGle 153
l+ + ++ le+ ++P+e++s+avfrcv+vs++d +e +aYqtav+igGhvfkGiLydqG++
Phvul.001G009800.1 212 LPIT--TTGLEVGQFPPELNSPAVFRCVKVSAMDAPDERYAYQTAVNIGGHVFKGILYDQGMD 272
6655..5679999************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF05142 | 3.8E-65 | 123 | 272 | IPR007818 | Protein of unknown function DUF702 |
| TIGRFAMs | TIGR01623 | 1.2E-26 | 125 | 167 | IPR006510 | Zinc finger, lateral root primordium type 1 |
| TIGRFAMs | TIGR01624 | 1.1E-24 | 223 | 271 | IPR006511 | Lateral Root Primordium type 1, C-terminal |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009299 | Biological Process | mRNA transcription | ||||
| GO:0048467 | Biological Process | gynoecium development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 334 aa Download sequence Send to blast |
MAGFFSLGGR HNKAEQEEDQ REDNNTNNTN HNNSQFLFRN VNEEIYNTNK GFEIWPQSSY 60 HHNFTNYYSF GVGPSRRNNA NTNNSSSNNV NDDVSVSFSD DSNRFGFTVM RSAGGVGGGG 120 GGGMNCQDCG NQAKKDCSHL RCRTCCKSRG FQCQTHVKST WVPAAKRRER QQQLATLQQQ 180 NQPPQFRGDH SKRHRESIEG AAAGGSLACA PLPITTTGLE VGQFPPELNS PAVFRCVKVS 240 AMDAPDERYA YQTAVNIGGH VFKGILYDQG MDGPYASAGC EGSSGGGGEA RALTLMAATT 300 TATTAGNPFD ASLYTAPMNA FMAGTQFFPP PRS* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds DNA on 5'-ACTCTAC-3' and promotes auxin homeostasis-regulating gene expression (e.g. YUC genes), as well as genes affecting stamen development, cell expansion and timing of flowering. Synergistically with other SHI-related proteins, regulates gynoecium, stamen and leaf development in a dose-dependent manner, controlling apical-basal patterning. Promotes style and stigma formation, and influences vascular development during gynoecium development. May also have a role in the formation and/or maintenance of the shoot apical meristem (SAM). {ECO:0000269|PubMed:16740146}. | |||||
| UniProt | Transcription activator that binds DNA on 5'-ACTCTAC-3' and promotes auxin homeostasis-regulating gene expression (e.g. YUC genes), as well as genes affecting stamen development, cell expansion and timing of flowering. Synergistically with other SHI-related proteins, regulates gynoecium, stamen and leaf development in a dose-dependent manner, controlling apical-basal patterning. Promotes style and stigma formation, and influences vascular development during gynoecium development. May also have a role in the formation and/or maintenance of the shoot apical meristem (SAM). Regulates anther dehiscence and floral development. {ECO:0000269|PubMed:16740146, ECO:0000269|PubMed:20706774}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00152 | DAP | Transfer from AT1G19790 | Download |
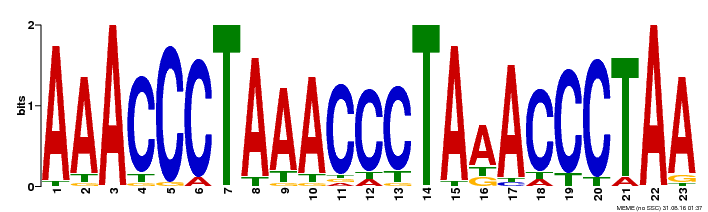
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Phvul.001G009800.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015036 | 1e-162 | AP015036.1 Vigna angularis var. angularis DNA, chromosome 3, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007160701.1 | 0.0 | hypothetical protein PHAVU_001G009800g | ||||
| Swissprot | Q9FXH7 | 7e-83 | SRS7_ARATH; Protein SHI RELATED SEQUENCE 7 | ||||
| Swissprot | Q9LQZ5 | 8e-83 | SRS5_ARATH; Protein SHI RELATED SEQUENCE 5 | ||||
| TrEMBL | V7CTW6 | 0.0 | V7CTW6_PHAVU; Uncharacterized protein | ||||
| STRING | XP_007160701.1 | 0.0 | (Phaseolus vulgaris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF882 | 34 | 125 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G75520.1 | 7e-74 | SHI-related sequence 5 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Phvul.001G009800.1 |
| Entrez Gene | 18640008 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



