 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.J694200.2.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 385aa MW: 42316.7 Da PI: 8.1052 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 88.9 | 4.8e-28 | 267 | 317 | 1 | 51 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQ 51
k rlrWt eLHerFveav++L G+ekAtPk++l+ mkv+gLt++hvkSHLQ
Pavir.J694200.2.p 267 KSRLRWTLELHERFVEAVNKLEGPEKATPKAVLKQMKVEGLTIYHVKSHLQ 317
68************************************************* PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 2.4E-25 | 265 | 317 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 1.33E-13 | 265 | 318 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 3.1E-21 | 267 | 318 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.0E-7 | 269 | 317 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 385 aa Download sequence Send to blast |
MSSQSVATGE QIIAPNQIVH ACTSTQTSVH KLFDAKLDHG FLIDDTLSST SQSSNIKTEL 60 IRTSSLSRSL SVNLQKRSPE SDPESPLSHV SHPKFSDPIL SNSSTFCTSL FSSSSKNTDP 120 CRQMGTLPFL PHPPKCEQQV SAGQSSSSSL LLSGDTGNAL DEAGQSDDLK DFLNLSGDAS 180 DGSFHGENNI LAFDEQMEFQ FLSEQLGIAI TDNEESPHLD DIYGTPPQLS SLPVSSCSNQ 240 SKQNLGSPVK VQLSSSRSSS DSATTNKSRL RWTLELHERF VEAVNKLEGP EKATPKAVLK 300 QMKVEGLTIY HVKSHLQVKV YSFFPLKNQI VRCNLHFPLH LSCRSIDLRN IFLSLKKTRR 360 LPLRIRKHNW VAAAVIPAKL RIYK* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 2e-21 | 267 | 317 | 2 | 52 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 2e-21 | 267 | 317 | 2 | 52 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 2e-21 | 267 | 317 | 2 | 52 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 2e-21 | 267 | 317 | 2 | 52 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 2e-21 | 267 | 317 | 2 | 52 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 2e-21 | 267 | 317 | 2 | 52 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 2e-21 | 267 | 317 | 2 | 52 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 2e-21 | 267 | 317 | 2 | 52 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Pvr.1761 | 0.0 | callus| leaf| root| stem | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Expressed throughout all stages of plant growth. Increased expression in leaves before the booting stage. {ECO:0000269|PubMed:26082401}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in the root cap and in the exodermis of the root, in the root tip of lateral roots, in the mesophyll cells of the leaf, in pollen, vascular cylinder of the anther and the veins of the lemma, palea and pistils, and in the xylem and phloem regions of large vascular bundles, small vascular bundles and diffuse vascular bundles in node I (PubMed:26082401). {ECO:0000269|PubMed:26082401}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in phosphate starvation signaling (PubMed:26082401). Binds to P1BS, an imperfect palindromic sequence 5'-GNATATNC-3', to promote the expression of inorganic phosphate (Pi) starvation-responsive genes (PubMed:26082401). Functionally redundant with PHR1 and PHR2 in regulating Pi starvation response and Pi homeostasis (PubMed:26082401). {ECO:0000269|PubMed:26082401}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00354 | DAP | Transfer from AT3G13040 | Download |
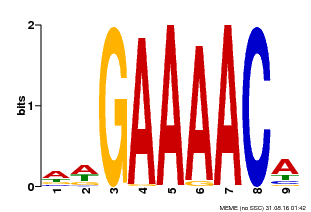
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.J694200.2.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated under Pi starvation conditions. {ECO:0000269|PubMed:26082401}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KF184668 | 1e-174 | KF184668.1 Saccharum hybrid cultivar R570 clone BAC 005I10 complete sequence. | |||
| GenBank | KF184707 | 1e-174 | KF184707.1 Saccharum hybrid cultivar R570 clone BAC 084G07 complete sequence. | |||
| GenBank | KF184833 | 1e-174 | KF184833.1 Saccharum hybrid cultivar R570 clone BAC 001I10 complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025812859.1 | 0.0 | protein PHOSPHATE STARVATION RESPONSE 3 | ||||
| Refseq | XP_025812860.1 | 0.0 | protein PHOSPHATE STARVATION RESPONSE 3 | ||||
| Refseq | XP_025812862.1 | 0.0 | protein PHOSPHATE STARVATION RESPONSE 3 | ||||
| Refseq | XP_025812863.1 | 0.0 | protein PHOSPHATE STARVATION RESPONSE 3 | ||||
| Swissprot | Q6YXZ4 | 1e-115 | PHR3_ORYSJ; Protein PHOSPHATE STARVATION RESPONSE 3 | ||||
| TrEMBL | A0A3L6PFL7 | 0.0 | A0A3L6PFL7_PANMI; Uncharacterized protein | ||||
| STRING | Pavir.J27649.1.p | 0.0 | (Panicum virgatum) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G13040.2 | 1e-49 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.J694200.2.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



