 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.J062800.2.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 791aa MW: 86692.4 Da PI: 6.2458 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 82.2 | 7e-26 | 1 | 53 | 26 | 78 |
HHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 26 vhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+h+ka++++v ++ qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+++
Pavir.J062800.2.p 1 MHAKASTAVVGNTVQRFCQQCSRFHLLQEFDEGKRSCRRRLAGHNRRRRKTRP 53
69************************************************875 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51141 | 21.939 | 1 | 51 | IPR004333 | Transcription factor, SBP-box |
| Gene3D | G3DSA:4.10.1100.10 | 1.7E-15 | 1 | 38 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 6.15E-23 | 1 | 55 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 2.2E-17 | 1 | 50 | IPR004333 | Transcription factor, SBP-box |
| CDD | cd00204 | 1.49E-9 | 547 | 677 | No hit | No description |
| SuperFamily | SSF48403 | 4.2E-8 | 571 | 677 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 0.34 | 572 | 601 | IPR002110 | Ankyrin repeat |
| Gene3D | G3DSA:1.25.40.20 | 5.3E-8 | 574 | 676 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 710 | 618 | 648 | IPR002110 | Ankyrin repeat |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 791 aa Download sequence Send to blast |
MHAKASTAVV GNTVQRFCQQ CSRFHLLQEF DEGKRSCRRR LAGHNRRRRK TRPDIAIGGT 60 ASIEDKVSNY LLLSLIGICA NLNSDSVQHS NSQELLSTLL KNLGSVAKSL EPKELCKLLE 120 AYQSLQNGSN AGTSGTTNAA EEAAGPSNSK LPFGNGSHRG QASSSVVPVQ SKATVVVTPE 180 PASCKLKDFD LNDTCNDMEG FEDGQEGSPT PAFKAADSPN CASWMQQDST QSPPQTSGNS 240 DSTSTQSLSS SNGDAQCRTD KIVFKLFNKV PSDLPPVLRS QILGWLSSSP TDIESYIRPG 300 CIILTVYLRL VESAWRELSG NMSLHLDKLL NSSTDNFWAS GLVFVMVRQQ LAFMHNGQIM 360 LDRPLAPSSH QYCKILRVRP VAAPYSATIN FRVEGFNLLS TSSRVICSYE GCCIFQEDTD 420 TVADDAEDIE CLSFCCSVPG SRGRGFIEVE DSGFSNGFFP FIIAEKDVCS EVSELESIFE 480 SSSDEYAVVD DNARDQALEF LNELGWLLHR ANRMSKQAET DTPLAAFNMW RFRKLGIFAM 540 EREWCAVVEM LLNFLFIGLV DLGSQSPEEM VLSENLLHAA VRRKSVKMVR FLLRYKPNKN 600 SKGTAQTYLF RPDALGPLTI TPLHIAAATS DAEDVLDALT DDPGLVGISA WSNARDETGF 660 TPEDYARQRG NDAYLNLVHK KIDKHLGKGH VVLGVPSSMC SVIPDGAKPG DVSLEICRPM 720 SASVPRCLLC TQQARVYPNS GARTFLYRPA MLTVMGVAVV CVCVGILLHT FPRVYAAPTF 780 RWELLERGPM * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 1e-17 | 1 | 50 | 35 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Pvr.835 | 0.0 | callus| leaf| root| stem | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Ubiquitous. {ECO:0000269|PubMed:16861571}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
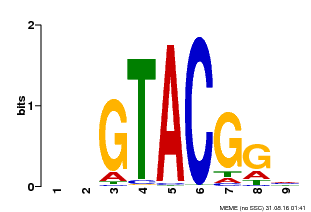
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.J062800.2.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT038852 | 0.0 | BT038852.1 Zea mays full-length cDNA clone ZM_BFb0336G12 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025793312.1 | 0.0 | squamosa promoter-binding-like protein 6 | ||||
| Swissprot | Q75LH6 | 0.0 | SPL6_ORYSJ; Squamosa promoter-binding-like protein 6 | ||||
| TrEMBL | A0A2S3IGN3 | 0.0 | A0A2S3IGN3_9POAL; Uncharacterized protein | ||||
| STRING | Pavir.J01500.1.p | 0.0 | (Panicum virgatum) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 1e-179 | squamosa promoter binding protein-like 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.J062800.2.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



