 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.9NG121500.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 317aa MW: 34823 Da PI: 6.1678 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 32.8 | 1.3e-10 | 147 | 195 | 3 | 55 |
HHHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 3 rahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
+h+ +Er RR++i +++ L++l+P + sk Ka +L + ++Y++sLq
Pavir.9NG121500.1.p 147 NSHSLAERLRRKKISERMKLLQDLVPGC----SKITGKAVMLDEIINYVQSLQ 195
58**************************....899*****************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF47459 | 1.44E-16 | 143 | 211 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 15.447 | 144 | 194 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 2.7E-16 | 147 | 211 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 5.3E-8 | 147 | 195 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 2.64E-11 | 148 | 199 | No hit | No description |
| SMART | SM00353 | 4.6E-10 | 150 | 200 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009637 | Biological Process | response to blue light | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 317 aa Download sequence Send to blast |
MYSGQQSDQC PSANSGREFS EANRNTVTMH QKMGYNSGPY GFGPYNMGLE ERPGLYQSSS 60 GTFSQNIQMS DEHSGGVKKR KGMDDCVAML QNAGDQQTEG SSQPERISLE GNRKISPKMQ 120 SKEDSSDGDG TKEDYVHVRA KQGQATNSHS LAERLRRKKI SERMKLLQDL VPGCSKITGK 180 AVMLDEIINY VQSLQRQVEF LSMKLATVNP ELGFDIEQIL SKQMMLSQDR HFAFYGVDPG 240 SSSLASQFSQ GIMQPPMMCN ISNPADVLQG TIHDVSTMNQ IPAMWEGLQN LPHMNFNPGV 300 AADSSANNSG SMKIEQ* |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed constitutively in roots, leaves, stems, and flowers. {ECO:0000269|PubMed:12679534}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator involved in cell elongation. Regulates the expression of a subset of genes involved in cell expansion by binding to the G-box motif. Binds to chromatin DNA of the FT gene and promotes its expression, and thus triggers flowering in response to blue light (PubMed:24130508). {ECO:0000269|PubMed:23161888, ECO:0000269|PubMed:24130508}. | |||||
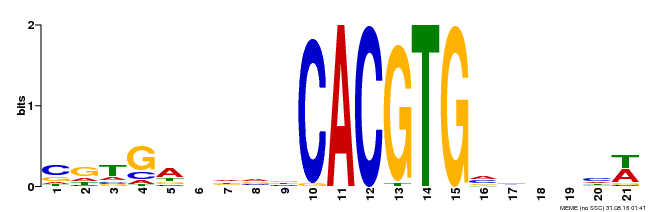
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00135 | DAP | Transfer from AT1G10120 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.9NG121500.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT035348 | 0.0 | BT035348.1 Zea mays full-length cDNA clone ZM_BFb0057A01 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025796141.1 | 0.0 | transcription factor bHLH74-like | ||||
| Swissprot | Q6NKN9 | 1e-51 | BH074_ARATH; Transcription factor bHLH74 | ||||
| TrEMBL | A0A2S3IIE7 | 0.0 | A0A2S3IIE7_9POAL; Uncharacterized protein | ||||
| TrEMBL | A0A2T7C1I4 | 0.0 | A0A2T7C1I4_9POAL; Uncharacterized protein | ||||
| TrEMBL | A0A3L6S4S4 | 0.0 | A0A3L6S4S4_PANMI; Transcription factor bHLH49-like | ||||
| STRING | Pavir.Ia00718.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6033 | 36 | 50 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G10120.1 | 6e-54 | bHLH family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.9NG121500.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



