 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.7NG427500.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 943aa MW: 104171 Da PI: 6.1734 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 70.5 | 2.2e-22 | 143 | 244 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkl 84
f+k lt sd++++g +++p++ a++ ++++++ ++l+++d++ W++++iyr++++r++lt+GW+ Fv +++Lk+gD+v+F
Pavir.7NG427500.1.p 143 FCKNLTASDTSTHGGFSVPRRAADKLfpqldySMQPPN-QELIVRDLHDSLWTFRHIYRGQPKRHLLTTGWSLFVGSKRLKAGDSVLFI- 230
89*********************999*****9555555.49************************************************. PP
-SSSEE..EEEEE-S CS
B3 85 dgrsefelvvkvfrk 99
++++ +l+v+v+r+
Pavir.7NG427500.1.p 231 -RDEKSQLLVGVRRA 244
.4577778*****97 PP
| |||||||
| 2 | Auxin_resp | 113.8 | 1.6e-37 | 269 | 351 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
aahaas++ F+++YnPr+s+s Fv+++ +++ka++++ svGmRf m+fete+ss+rr++Gt+vg+sd+dp+rWpnSkWr+L+
Pavir.7NG427500.1.p 269 AAHAASSGGSFTIYYNPRTSPSPFVIPLTRYNKATYMQPSVGMRFAMMFETEESSKRRCTGTIVGISDYDPMRWPNSKWRNLQ 351
79*******************************************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 9.16E-44 | 133 | 272 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 2.8E-39 | 136 | 257 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 8.67E-20 | 142 | 243 | No hit | No description |
| SMART | SM01019 | 2.6E-21 | 143 | 245 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 12.492 | 143 | 245 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 5.2E-20 | 143 | 244 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 3.9E-33 | 269 | 351 | IPR010525 | Auxin response factor |
| Pfam | PF02309 | 1.7E-9 | 838 | 926 | IPR033389 | AUX/IAA domain |
| PROSITE profile | PS51745 | 24.119 | 839 | 923 | IPR000270 | PB1 domain |
| SuperFamily | SSF54277 | 3.79E-8 | 857 | 918 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009908 | Biological Process | flower development | ||||
| GO:0009942 | Biological Process | longitudinal axis specification | ||||
| GO:0010305 | Biological Process | leaf vascular tissue pattern formation | ||||
| GO:0048364 | Biological Process | root development | ||||
| GO:0048507 | Biological Process | meristem development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016020 | Cellular Component | membrane | ||||
| GO:0042802 | Molecular Function | identical protein binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 943 aa Download sequence Send to blast |
MASSQEKATS GVLRNAAALL DEMQLMGEAQ GAKKVINSEL WHACAGPLVC LPQRGSLVYY 60 FPQGHSEQVA ATTKKVPNSR IPNYPSLPSQ LLCQVHNITL HADKETDEIY AQMTLQPVHS 120 ETDVFPIPTL GAYTKSKHPT EYFCKNLTAS DTSTHGGFSV PRRAADKLFP QLDYSMQPPN 180 QELIVRDLHD SLWTFRHIYR GQPKRHLLTT GWSLFVGSKR LKAGDSVLFI RDEKSQLLVG 240 VRRATRQQPP LSSSVLSTDS MHIGVLAAAA HAASSGGSFT IYYNPRTSPS PFVIPLTRYN 300 KATYMQPSVG MRFAMMFETE ESSKRRCTGT IVGISDYDPM RWPNSKWRNL QVEWDEHGYG 360 ERPERVNLWD IETPENSLVF SSPLNSKRQC LPSYGVPGLQ IDNLSSIPRA QANPFGNLQH 420 MPGIGSELAL MLLNQPGQNL GSPLACQQSS FSSIIQNVKH GYIPPLTFGS SNGSIKQENM 480 ISNEAQQQLN APNIEKDDQV SCEVQPSIDS ISEQELNVKA RGPRNTDSYS SQSISDQNSK 540 GEPRTKTRRS KKGLSHKSIS DKSELSSVPS QIYDNQRHCS EPKLVDCEAE QVNCEDSSGA 600 LTRGGFAGAQ VQHVEQHELL SPAKLDSSKS PDGGKSVSSF PNQGCSPQFF EGLDWVIQPS 660 YYQDSNGIHS VSGSENIFNQ STDITSTINA DTMEAFQTSC LSECFPNSVQ EFISSPDLNS 720 LTFMSHDMQH LDGQHEVNNL PSTSNSYVQM TYSEESGNQS ASLSGLHMEA IHINSSCSEP 780 LATGSFDTGM FSKLPDLRES QVLPLHEIHN SSMGKPSCSM EAAEYSIDRS AKPMKPPVRT 840 FTKVQKLGSV GRSIDVTRYR DYHELRSAIA CMFGLQGKLE HPGSSDWKLV YVDYENDVLL 900 VGDDPWEEFI NCVRCIRILS PSEVQQMSEN GVHVLNDCIQ IA* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldu_A | 0.0 | 17 | 374 | 31 | 390 | Auxin response factor 5 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Pvr.26057 | 0.0 | callus | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, culms, leaves and young panicles. {ECO:0000269|PubMed:17408882}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00153 | DAP | Transfer from AT1G19850 | Download |
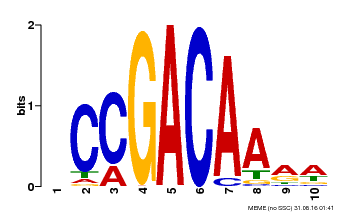
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.7NG427500.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025824429.1 | 0.0 | auxin response factor 11 | ||||
| Swissprot | Q01K26 | 0.0 | ARFK_ORYSI; Auxin response factor 11 | ||||
| Swissprot | Q8S983 | 0.0 | ARFK_ORYSJ; Auxin response factor 11 | ||||
| TrEMBL | A0A2S3IBD3 | 0.0 | A0A2S3IBD3_9POAL; Auxin response factor | ||||
| STRING | Pavir.Gb00117.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP8673 | 32 | 41 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G19850.1 | 0.0 | ARF family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.7NG427500.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



