 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.7KG316000.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 448aa MW: 49028.9 Da PI: 8.2148 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 52.3 | 1.3e-16 | 54 | 98 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT++E++l+++a +q+G W++I +++g +t+ q++s+ qk+
Pavir.7KG316000.1.p 54 REKWTEDEHKLFLEALQQHGRA-WRRIQEHIG-SKTAVQIRSHAQKF 98
789*****************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 3.21E-15 | 48 | 101 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 20.136 | 49 | 103 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.0E-9 | 51 | 104 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.6E-15 | 52 | 101 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 3.5E-13 | 53 | 101 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 4.2E-14 | 54 | 97 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.38E-10 | 56 | 99 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 448 aa Download sequence Send to blast |
MARFQEIKAT NDRKPAADRI GHADFMDHPK KTLDSPEGPR APKARKPYTI SKQREKWTED 60 EHKLFLEALQ QHGRAWRRIQ EHIGSKTAVQ IRSHAQKFFS KVIRESSGDS NSIAAPPQIY 120 IPPARPKRKP AHPYPRNLGN SLCKDASTIK QLEKPQLKIQ LLSEQENCSP KSVLTTAQID 180 SETMATEGSG SPASSVYMEE KCLTPSTSVG ELGVQVALSK DATTSNGAAC GIPEGPVLRL 240 FGKRVVVNNL HQQPNSDTGN LKHAADMELD ASAETPTSGT GKFSSHGAEE GRTWSPWLTG 300 TQQFMYYLPQ GEVLSVHSAC QFLSYSNVSI SYGVLNPQTV ASKKQQHQPS QASDCKFTKA 360 EGSWAESITT SSSVPETTQN SDSIESSQVN NDDDEVILVP GSRKYLSTVP TYLRGFVPYK 420 KCTAQSKMPQ SHVPGEEADG DMTRLCL* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Pvr.34707 | 0.0 | callus| leaf| root | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00515 | DAP | Transfer from AT5G17300 | Download |
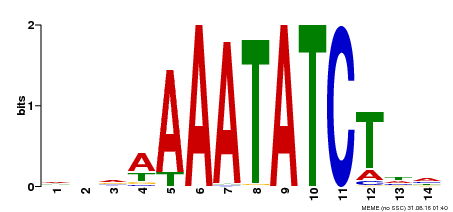
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.7KG316000.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025824173.1 | 0.0 | uncharacterized protein LOC112899782 | ||||
| TrEMBL | A0A2S3I9P7 | 0.0 | A0A2S3I9P7_9POAL; Uncharacterized protein | ||||
| STRING | Pavir.Gb00859.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2722 | 36 | 86 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G17300.1 | 2e-32 | MYB_related family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.7KG316000.1.p |



