 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.6KG307800.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 352aa MW: 36953.1 Da PI: 8.4252 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 44.5 | 3.4e-14 | 273 | 324 | 5 | 56 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkkev 56
+r+rr++kNRe+A rsR+RK+a++ eLe +v++L++ N +L+k+ ee+ ++
Pavir.6KG307800.1.p 273 RRQRRMIKNRESAARSRARKQAYTMELEAEVQKLKELNAELQKKQEEMMEMQ 324
79****************************************9999998875 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 8.5E-13 | 269 | 333 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 10.932 | 271 | 322 | IPR004827 | Basic-leucine zipper domain |
| CDD | cd14707 | 5.83E-28 | 273 | 327 | No hit | No description |
| Pfam | PF00170 | 1.5E-12 | 273 | 325 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 1.69E-10 | 273 | 322 | No hit | No description |
| Gene3D | G3DSA:1.20.5.170 | 1.3E-14 | 273 | 322 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 276 | 291 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009738 | Biological Process | abscisic acid-activated signaling pathway | ||||
| GO:0010255 | Biological Process | glucose mediated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0000976 | Molecular Function | transcription regulatory region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 352 aa Download sequence Send to blast |
MDLKDGWGRG AAGPGPLSRQ GSIYSLTFDE LQSTLGGMGG GLGKDFGSMN MDELLRSIWT 60 AEESQVMASA SAAGAPAEDG AAALQRQGSL TLPRTLSVKT VDEVWRDFVR EVPPPAGAEP 120 QPTRQPTLGE MTLEEFLVRA GVVRDNPAAA AMAAAAAVPA QPVAPRPIQA VSNGASIFFG 180 NFGAASDAGP GAMGFAPVGI GDQGMGNGLM PGVAGMASAA VTVSPVDTSV AQLDSVGKGN 240 GDLSSPMAPV PYPFVGVIRG RRSGAGVEKV VERRQRRMIK NRESAARSRA RKQAYTMELE 300 AEVQKLKELN AELQKKQEEM MEMQKNQVLE VVSNPYAQKK RCLQRTLTGP W* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Pvr.22531 | 0.0 | leaf| root | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, leaves and embryos. {ECO:0000269|PubMed:10611387}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that mediates abscisic acid (ABA) signaling. Binds specifically to the ABA-responsive element (ABRE) of the EMP1 and RAB16A gene promoters. {ECO:0000269|PubMed:10611387}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00186 | DAP | Transfer from AT1G45249 | Download |
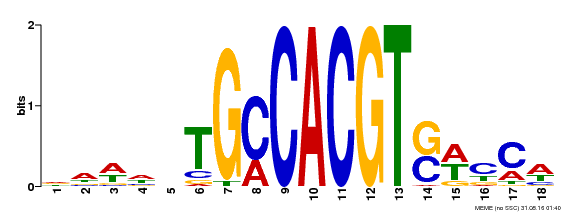
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.6KG307800.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By abscisic acid (ABA). {ECO:0000269|PubMed:10611387}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004973666.1 | 0.0 | bZIP transcription factor TRAB1 | ||||
| Swissprot | Q6ZDF3 | 1e-150 | TRAB1_ORYSJ; bZIP transcription factor TRAB1 | ||||
| TrEMBL | K3YIB0 | 0.0 | K3YIB0_SETIT; Uncharacterized protein | ||||
| STRING | Pavir.Fa00674.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP706 | 38 | 147 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G41070.3 | 2e-38 | bZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.6KG307800.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



