 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.6KG273500.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 294aa MW: 31082.1 Da PI: 9.2207 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 55.2 | 1.6e-17 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT+eEd+ lv +v+++G+g+W++++ g+ R+ k+c++rw +yl
Pavir.6KG273500.1.p 14 KGPWTPEEDLVLVSYVQEHGPGNWRAVPTNTGLMRCSKSCRLRWTNYL 61
79********************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 47.5 | 4e-15 | 67 | 112 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg+++ E++l++++ ++lG++ W++Ia++++ Rt++++k++w+++l
Pavir.6KG273500.1.p 67 RGNFSVKEEKLIIHLQALLGNR-WAAIASYLP-DRTDNDIKNYWNTHL 112
89********************.*********.************996 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 2.2E-24 | 5 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 24.975 | 9 | 65 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.38E-30 | 11 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 4.0E-13 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.2E-16 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.24E-11 | 16 | 61 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 1.1E-23 | 65 | 116 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 19.983 | 66 | 116 | IPR017930 | Myb domain |
| SMART | SM00717 | 8.0E-16 | 66 | 114 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.8E-14 | 67 | 112 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 7.11E-12 | 69 | 112 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0010468 | Biological Process | regulation of gene expression | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 294 aa Download sequence Send to blast |
MGRPPCCEKA GVKKGPWTPE EDLVLVSYVQ EHGPGNWRAV PTNTGLMRCS KSCRLRWTNY 60 LRPGIKRGNF SVKEEKLIIH LQALLGNRWA AIASYLPDRT DNDIKNYWNT HLKKKLATMG 120 ADGAGAGGAE ARSGRAAAPK GQWERRLQTD IHTARQALRE ALSLDPVKLP PAKTEPLPQP 180 APAQPSQAAY ASSAENIARL LEGWMRPGTG SAGTKAPSGS RSSASAVSGG EGASASHSGT 240 ARTPEGSTGT SKAEDAGPMP AFSMLESWLL DDGMGHGDAG LMGVPLADPC EFF* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1gv2_A | 5e-25 | 14 | 116 | 4 | 105 | MYB PROTO-ONCOGENE PROTEIN |
| Search in ModeBase | ||||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Expression in flowers increases as the flowers develop. {ECO:0000269|PubMed:1840903}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in flowers, leaves and weakly in seed pods. {ECO:0000269|PubMed:1840903}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor. {ECO:0000305}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00579 | DAP | Transfer from AT5G62470 | Download |
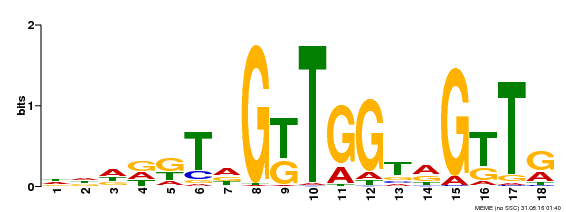
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.6KG273500.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT035749 | 1e-149 | BT035749.2 Zea mays full-length cDNA clone ZM_BFb0081I17 mRNA, complete cds. | |||
| GenBank | KJ727881 | 1e-149 | KJ727881.1 Zea mays clone pUT5974 MYB transcription factor (MYB83) mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025821217.1 | 1e-156 | myb-related protein 306-like isoform X1 | ||||
| Swissprot | P81392 | 1e-105 | MYB06_ANTMA; Myb-related protein 306 | ||||
| TrEMBL | A0A3L6PM73 | 1e-169 | A0A3L6PM73_PANMI; Uncharacterized protein | ||||
| STRING | Pavir.Fa00895.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP229 | 38 | 296 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G62470.2 | 3e-89 | myb domain protein 96 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.6KG273500.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



