 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.5NG613200.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 810aa MW: 90732.6 Da PI: 6.5164 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 73.3 | 3e-23 | 129 | 230 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFk 83
f+k+lt sd++++g +++ +++a+e+ +++++ ++l+ +d++g +W++++i+r++++r++l++GW+ Fv++++L +gD ++F
Pavir.5NG613200.1.p 129 FCKTLTASDTSTHGGFSVLRRHADEClppldmtQSPPT--QELVAKDLHGMEWRFRHIFRGQPRRHLLQSGWSVFVSSKRLVAGDAFIFL 216
99*****************************7555544..58************************************************ PP
E-SSSEE..EEEEE-S CS
B3 84 ldgrsefelvvkvfrk 99
+ +++el+v+v+r+
Pavir.5NG613200.1.p 217 --RGENGELRVGVRRA 230
..449999*****996 PP
| |||||||
| 2 | Auxin_resp | 112.4 | 4.3e-37 | 255 | 336 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
a+ha++tks+F+v+Y+Pr+s+seF++++++++++lk+++s+GmRf+m+fe+e+++e+r++Gt+vg ++ldp Wp+S Wr Lk
Pavir.5NG613200.1.p 255 AWHAINTKSMFTVYYKPRTSPSEFIIPYDQYMESLKNNYSIGMRFRMRFEGEEAPEQRFTGTIVGCENLDP-LWPDSSWRYLK 336
89*********************************************************************.9*******996 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 3.1E-40 | 120 | 244 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 4.45E-39 | 122 | 258 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 3.88E-21 | 127 | 229 | No hit | No description |
| SMART | SM01019 | 1.8E-23 | 129 | 231 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 4.0E-21 | 129 | 230 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 11.928 | 129 | 231 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 1.7E-33 | 255 | 336 | IPR010525 | Auxin response factor |
| Pfam | PF02309 | 1.9E-6 | 677 | 733 | IPR033389 | AUX/IAA domain |
| SuperFamily | SSF54277 | 4.76E-13 | 691 | 774 | No hit | No description |
| PROSITE profile | PS51745 | 24.865 | 697 | 790 | IPR000270 | PB1 domain |
| Pfam | PF02309 | 4.4E-8 | 743 | 782 | IPR033389 | AUX/IAA domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008285 | Biological Process | negative regulation of cell proliferation | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0010227 | Biological Process | floral organ abscission | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 810 aa Download sequence Send to blast |
MPPAAMAPPP PPQAPSNSGD PLYPELWRAC AGPLVTVPRP GDLVFYFPQG HIEQVEASMN 60 QVAGNQMRLY DLPSKLLCRV LNVELKAETE TDEVYAQIML MPEPEQNEVP SEKASSGAAA 120 TPRPAVRSFC KTLTASDTST HGGFSVLRRH ADECLPPLDM TQSPPTQELV AKDLHGMEWR 180 FRHIFRGQPR RHLLQSGWSV FVSSKRLVAG DAFIFLRGEN GELRVGVRRA MRQLCNVPSS 240 VISSQSMHLG VLATAWHAIN TKSMFTVYYK PRTSPSEFII PYDQYMESLK NNYSIGMRFR 300 MRFEGEEAPE QRFTGTIVGC ENLDPLWPDS SWRYLKVRWD EPSTIPRPDR VSPWKIEPAS 360 SPPVNPLPLS SRVKRPRQAP PPSPESSVLT KEGATKIDIS SAQTQHQNSV LRGQEQMTLR 420 NNMTESTDSD ATVQKPMVWS PSPNGKTHTS FQQRPSMDNW MPLGRRENDF KDTRSAFKDA 480 RTSSQSYGDT QGFFMPTFDE NHHHLSFNNQ FQDQGSAHHF ADPYFYMPQQ PSLTVEPSTR 540 TQSASNELRF WSDQNTVYGN TNDQQKGFTF AQNPSSWLNQ PFPHAEQPRV VRPHATVAAF 600 DLEKTREGSG FKIFGFKVDT ASASPIQLSS PMSAMREHVV QTQPSASVNE LQHVQTECLP 660 EGSVSTAGTA TENEKSNQQA PQSSKDIQSK SQGASTRSCT KVHKQGVALG RSVDLSKFSD 720 YDELKAELDK MFEFEGELVS ANKNWQIVYT DNEGDMMLVG DDPWEEFCSI VRKIYIYTKE 780 EVQKMNSKSS ALRKEEPLAS GEVCAATNE* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 1e-166 | 20 | 361 | 17 | 357 | Auxin response factor 1 |
| 4ldw_A | 1e-166 | 20 | 361 | 17 | 357 | Auxin response factor 1 |
| 4ldw_B | 1e-166 | 20 | 361 | 17 | 357 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Pvr.346 | 0.0 | callus| leaf| root| stem | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, culms, leaves and young panicles. {ECO:0000269|PubMed:17408882}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
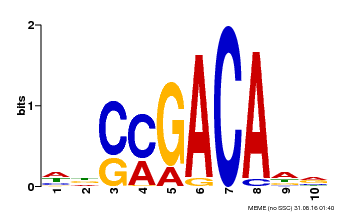
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00574 | DAP | Transfer from AT5G62000 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.5NG613200.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025815802.1 | 0.0 | auxin response factor 4-like | ||||
| Swissprot | Q5JK20 | 0.0 | ARFD_ORYSJ; Auxin response factor 4 | ||||
| TrEMBL | A0A2T7DC64 | 0.0 | A0A2T7DC64_9POAL; Auxin response factor | ||||
| STRING | Pavir.Eb03734.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1178 | 36 | 111 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G62000.3 | 0.0 | auxin response factor 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.5NG613200.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



