 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.5NG548200.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 599aa MW: 65031.4 Da PI: 5.7659 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 52.4 | 1.3e-16 | 89 | 136 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT Ed +lvd+vk++G g+W+++ + g+ R++k+c++rw ++l
Pavir.5NG548200.1.p 89 KGPWTSAEDAILVDYVKKHGEGNWNAVQKNTGLFRCGKSCRLRWANHL 136
79******************************************9986 PP
| |||||||
| 2 | Myb_DNA-binding | 52.5 | 1.1e-16 | 142 | 185 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
+g++T+eE+ l++++++++G++ W++ a++++ gRt++++k++w++
Pavir.5NG548200.1.p 142 KGAFTPEEERLIIQLHAKMGNK-WARMAAHLP-GRTDNEIKNYWNT 185
799*******************.*********.***********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF001693 | 5.2E-255 | 46 | 598 | IPR016310 | Transcription factor, GAMYB |
| PROSITE profile | PS51294 | 16.625 | 84 | 136 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 5.24E-31 | 86 | 183 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 2.1E-14 | 88 | 138 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.2E-14 | 89 | 136 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.3E-23 | 90 | 143 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 3.08E-11 | 91 | 136 | No hit | No description |
| PROSITE profile | PS51294 | 25.656 | 137 | 191 | IPR017930 | Myb domain |
| SMART | SM00717 | 2.1E-17 | 141 | 189 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 7.6E-16 | 142 | 185 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 8.17E-13 | 144 | 185 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 3.6E-26 | 144 | 189 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009789 | Biological Process | positive regulation of abscisic acid-activated signaling pathway | ||||
| GO:0043068 | Biological Process | positive regulation of programmed cell death | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0045926 | Biological Process | negative regulation of growth | ||||
| GO:0048235 | Biological Process | pollen sperm cell differentiation | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 599 aa Download sequence Send to blast |
MLHGLPPRPL SARNQAGRRY MNQGDEEENR GRRKRGRMRW RPDRAAMYRV KSESDCEMML 60 QDQMDSPVAD DGSSGGGSPH RGSGPPLKKG PWTSAEDAIL VDYVKKHGEG NWNAVQKNTG 120 LFRCGKSCRL RWANHLRPNL KKGAFTPEEE RLIIQLHAKM GNKWARMAAH LPGRTDNEIK 180 NYWNTRIKRC QRAGLPIYPA SVCNQSSSED QQISGDFNCG ENISNDLLSG NSLYLPDFTS 240 DNFIANTEAL PYAPQLSAAS ISNLLGQSFA SKNCSFMDQA DQSGMLKQSG YVLPALSDTV 300 DGVLSSVDQF SNDSEKLKQA LGFDYLNEAN ASSKSITPFG VTLSGSHAFL NGIFSASRPI 360 NGPLKMELPS LQDTESDPNS WLKYTLAPAM QPTELVDPYL HSPIATPSVK SESASPRNSG 420 LLEELLHEAQ ALRSGKNQLP SVRSSSSSAG TPCETTTVVS PEFDLCQEYL DEHHSSFLNE 480 CAPFSGYSFT ESTPVSAASP DIFQLSKISP AQSPSMGSGE QAVEPKHDTA GSPHPENLRP 540 DALFSGNTAD PSSFNDAIAM LLGNDMNAEP KPVLGDGIAF GSSSWGNMPH ACEISEFK* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 5e-32 | 85 | 189 | 3 | 106 | B-MYB |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Pvr.22992 | 0.0 | leaf | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator of gibberellin-dependent alpha-amylase expression in aleurone cells. Involved in pollen and floral organs development. May bind to the 5'-TAACAAA-3' box of alpha-amylase promoter. {ECO:0000269|PubMed:9150608}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00490 | DAP | Transfer from AT5G06100 | Download |
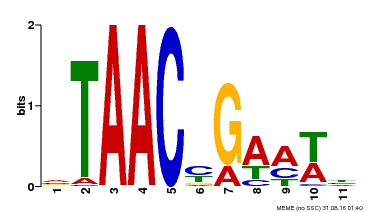
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.5NG548200.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By gibberellin in aleurone cells. {ECO:0000269|PubMed:9150608}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT033925 | 0.0 | BT033925.1 Zea mays full-length cDNA clone ZM_BFc0044D10 mRNA, complete cds. | |||
| GenBank | BT062692 | 0.0 | BT062692.1 Zea mays full-length cDNA clone ZM_BFb0334D20 mRNA, complete cds. | |||
| GenBank | BT063828 | 0.0 | BT063828.1 Zea mays full-length cDNA clone ZM_BFc0110K24 mRNA, complete cds. | |||
| GenBank | HQ858750 | 0.0 | HQ858750.1 Zea mays clone UT1459 MYBGA transcription factor mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025819835.1 | 0.0 | transcription factor GAMYB-like | ||||
| Swissprot | A2WW87 | 0.0 | GAM1_ORYSI; Transcription factor GAMYB | ||||
| TrEMBL | A0A2S3HQQ9 | 0.0 | A0A2S3HQQ9_9POAL; Transcription factor | ||||
| TrEMBL | A0A2T7DFC7 | 0.0 | A0A2T7DFC7_9POAL; Transcription factor | ||||
| STRING | Pavir.Ea03206.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3947 | 36 | 64 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G06100.3 | 4e-87 | myb domain protein 33 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.5NG548200.1.p |



