 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.5NG543900.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 449aa MW: 49347.4 Da PI: 7.0373 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 29.7 | 1.4e-09 | 163 | 203 | 5 | 45 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaL 45
k rr+++NReAAr+sR+RKka+i++Le +L++ ++L
Pavir.5NG543900.1.p 163 KTLRRLAQNREAARKSRLRKKAYIQNLESSRLKLTQLEQEL 203
7889************************9777777666555 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 3.3E-8 | 159 | 222 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 9.312 | 161 | 205 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 4.25E-7 | 163 | 206 | No hit | No description |
| CDD | cd14708 | 3.29E-23 | 163 | 215 | No hit | No description |
| Pfam | PF00170 | 9.4E-7 | 163 | 203 | IPR004827 | Basic-leucine zipper domain |
| Gene3D | G3DSA:1.20.5.170 | 1.2E-7 | 164 | 207 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 166 | 181 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF14144 | 6.7E-33 | 245 | 320 | IPR025422 | Transcription factor TGA like domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009410 | Biological Process | response to xenobiotic stimulus | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005737 | Cellular Component | cytoplasm | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 449 aa Download sequence Send to blast |
MRSFGPPQHS IHTDMNSMQP SQVTNFGALA QSAGFRIEDL ANLNTSTLFN LKTNTINNNP 60 LQFGSYGKPI SPSHINTTTT ATAAVRIDPQ SLEHQTGAQQ NLVALPTGNI ENWGESAMAD 120 SPMIDTSTDP DTDERNQMFE QGLVAGPTAS DSSDKSRDKL DQKTLRRLAQ NREAARKSRL 180 RKKAYIQNLE SSRLKLTQLE QELQRARQQG IFISTSGDQP QSANGNGALA FDMEYARWLE 240 EHNKHVNELR AAVNAHAGDN DLRGIVDSIM AHYDEIFRLK GVAAKADVFH VLSGMWKTPA 300 ERCFMWLGGF RSSELLKLLA GQLEPLTDQQ LVGISNLQQS SQQAEDALSQ GMEALQQSLA 360 ETLASGSLGP AGSSGNVANY MGQMAMAMGK LGTLENFLRQ ADNLRLQTLQ QMQRILTTRQ 420 SARALLAISD YFSRLRALSS LWLARPRE* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Pvr.8649 | 0.0 | callus| leaf| stem | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional regulator involved in defense response. {ECO:0000250|UniProtKB:Q7X993}. | |||||
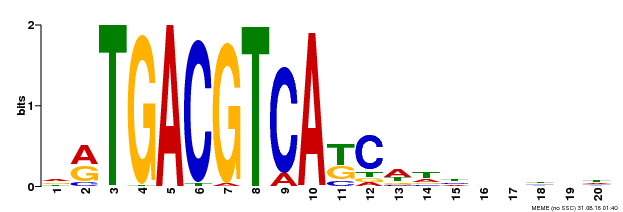
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00492 | DAP | Transfer from AT5G06950 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.5NG543900.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT034456 | 0.0 | BT034456.1 Zea mays full-length cDNA clone ZM_BFc0173A06 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025816950.1 | 0.0 | transcription factor TGAL1 isoform X1 | ||||
| Swissprot | Q6IVC2 | 0.0 | TGAL1_ORYSJ; Transcription factor TGAL1 | ||||
| TrEMBL | A0A2S3HR67 | 0.0 | A0A2S3HR67_9POAL; Uncharacterized protein | ||||
| TrEMBL | A0A2T7DFG0 | 0.0 | A0A2T7DFG0_9POAL; Uncharacterized protein | ||||
| STRING | Sb03g037560.1 | 0.0 | (Sorghum bicolor) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP486 | 38 | 185 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G12250.2 | 1e-142 | TGACG motif-binding factor 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.5NG543900.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



