 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.5NG136300.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 812aa MW: 86864 Da PI: 5.1508 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 69.3 | 4.8e-22 | 104 | 159 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r eL+++lgL+ rqVk+WFqNrR+++k
Pavir.5NG136300.1.p 104 KKRYHRHTPQQIQELEALFKECPHPDEKQRGELSRRLGLDPRQVKFWFQNRRTQMK 159
678899***********************************************999 PP
| |||||||
| 2 | START | 183.7 | 1e-57 | 322 | 553 | 2 | 206 |
HHHHHHHHHHHHHHHC-TT-EEEE......EXCCTTEEEEEEESSS......SCEEEEEEEECCS.CHHHHHHHHHCCCGGCT-TT-S.. CS
START 2 laeeaaqelvkkalaeepgWvkss......esengdevlqkfeeskv.....dsgealrasgvvd.mvlallveellddkeqWdetla.. 77
la a++elvk+a+ +ep+W s e++n +e+ +f + + + +ea+r+sg+v+ ++++ lve+l+d + +W+ ++
Pavir.5NG136300.1.p 322 LAINAMDELVKLAQIDEPLWLPSLngspnkEMLNFEEYAHTFLPCIGvkpmgYVSEASRESGLVIiDDSVALVETLMDER-RWSDMFScm 410
78899*******************999999999999999999998889**************997245679*********.********* PP
..EEEEEEEECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--......-TTSEE-EESSEEEE CS
START 78 ..kaetlevissg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe.....sssvvRaellpSgil 153
ka+++e ++sg gal +m+aelq+lsplvp R+++f+R+++ql +g w++vdvS+d ++ ++ vR+ +lpSg++
Pavir.5NG136300.1.p 411 iaKATVIEEVTSGiagsrnGALLVMKAELQVLSPLVPiREVTFLRFCKQLAEGAWAVVDVSIDGLLRDQNsattsNAANVRCMRLPSGCV 500
**************************************************************85322222234568999*********** PP
EEEECTCEEEEEEEE-EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 154 iepksnghskvtwvehvdlkgrlphwllrslvksglaegaktwvatlqrqcek 206
+++++ng +kvtwveh++++++++h+l+r+l++sgla+ga++w+a lqrqce+
Pavir.5NG136300.1.p 501 MQDTPNGFCKVTWVEHTEYDEASVHQLYRPLLRSGLAFGARRWLAMLQRQCEC 553
***************************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 7.52E-22 | 85 | 161 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 7.6E-24 | 89 | 161 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 18.285 | 101 | 161 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 4.8E-21 | 102 | 165 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 6.90E-20 | 103 | 161 | No hit | No description |
| Pfam | PF00046 | 1.5E-19 | 104 | 159 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 136 | 159 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 45.75 | 312 | 556 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 4.18E-30 | 315 | 553 | No hit | No description |
| CDD | cd08875 | 1.72E-112 | 316 | 552 | No hit | No description |
| SMART | SM00234 | 3.3E-46 | 321 | 553 | IPR002913 | START domain |
| Pfam | PF01852 | 2.5E-50 | 322 | 553 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.65E-21 | 573 | 776 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 812 aa Download sequence Send to blast |
MSFGSLFDGG AGGGSGGGMQ FPFSAGFSSS PGLSLGLDNN AAAGGGMGGG RALPGGPGAG 60 GGSGAARDAD AENDSRSGSD HLDAMSGGGE DEDDAEPGNP RKRKKRYHRH TPQQIQELEA 120 LFKECPHPDE KQRGELSRRL GLDPRQVKFW FQNRRTQMKT QLERHENALL KQENDKLRAE 180 NMTIREAMHS PMCGSCGSPA MLGEVSLEEQ HLCIENARLK DELNRVYALA TKFLGKPMSM 240 LAGPLMQPHL SSLPMPSSSL ELAIGDFRGL GSMPSATMPG SMGEFAGGVS SPLGTVITPA 300 RATGSAPLAM VRIDDRSMLL ELAINAMDEL VKLAQIDEPL WLPSLNGSPN KEMLNFEEYA 360 HTFLPCIGVK PMGYVSEASR ESGLVIIDDS VALVETLMDE RRWSDMFSCM IAKATVIEEV 420 TSGIAGSRNG ALLVMKAELQ VLSPLVPIRE VTFLRFCKQL AEGAWAVVDV SIDGLLRDQN 480 SATTSNAANV RCMRLPSGCV MQDTPNGFCK VTWVEHTEYD EASVHQLYRP LLRSGLAFGA 540 RRWLAMLQRQ CECLAILMSP DTVSANDSSV ITQEGKRSML KLARRMTENF CAGVSASSAR 600 EWSKLDGATG SIEEDVRVMA RKSVDEPGEP PGVVLSAATS VWVPVAPEKL FNFLRDEQLR 660 AEWDILSNGG PMQEMANIAK GQEHGNSVSL LRASAMSANQ SSMLILQETC TDASGSMVVY 720 APVDIPAMQL VMNGGDSTYV ALLPSGFAIL PDGPSATTGH KTGGSLLTVA FQILVNSQPT 780 AKLTVESVVT VNNLISCTIK KIKTALQCDA T* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 99 | 105 | PRKRKKR |
| 2 | 100 | 104 | RKRKK |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Pvr.16640 | 0.0 | stem | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
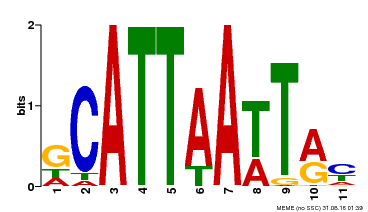
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.5NG136300.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT063249 | 0.0 | BT063249.1 Zea mays full-length cDNA clone ZM_BFc0042P14 mRNA, complete cds. | |||
| GenBank | BT064453 | 0.0 | BT064453.1 Zea mays full-length cDNA clone ZM_BFc0170E20 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025816451.1 | 0.0 | homeobox-leucine zipper protein ROC5-like | ||||
| Swissprot | Q6EPF0 | 0.0 | ROC5_ORYSJ; Homeobox-leucine zipper protein ROC5 | ||||
| TrEMBL | A0A2S3HX75 | 0.0 | A0A2S3HX75_9POAL; Uncharacterized protein | ||||
| STRING | Pavir.Eb00587.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1203 | 37 | 116 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G61150.1 | 0.0 | homeodomain GLABROUS 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.5NG136300.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



