 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.5NG037700.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 437aa MW: 49090.8 Da PI: 7.631 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 114.5 | 1.1e-35 | 44 | 186 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkk......leleevikevd....iykvePwdLpkkvkaeekewyfFskrdkkyatgkrknra... 77
lp+G++F+Ptd+el+ e+L++kv+++ +++ +i+++d i++++P++Lp v+ + + +fF++ +k+y+tg+rk+r+
Pavir.5NG037700.1.p 44 LPAGVKFDPTDQELI-EHLEAKVKDEGsrshplIDE--FIPTIDgedgICYTHPEKLP-GVTRDGLSKHFFHRPSKAYTTGTRKRRKiqt 129
799************.99******999677544333..3444433445**********.555666799******************7566 PP
NAM 78 .....tksgyWkatgkdkevlskkgelvglkktLvfy..kgrapkgektdWvmheyrl 128
+ +W++tgk+++v++ +g+++g+kk+Lv+y g+++k+ekt+Wvmh+y+l
Pavir.5NG037700.1.p 130 ecdvhKGETRWHKTGKTRPVMV-NGRQKGCKKILVLYtnFGKHRKPEKTNWVMHQYHL 186
6664433778************.999***********5547999999*********98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 2.09E-39 | 39 | 205 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 34.126 | 44 | 205 | IPR003441 | NAC domain |
| Pfam | PF02365 | 3.0E-16 | 45 | 186 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 437 aa Download sequence Send to blast |
MNKGHSSSSS KLINEKLEEH RISTAKHCPH CGEKIDSKPD WVGLPAGVKF DPTDQELIEH 60 LEAKVKDEGS RSHPLIDEFI PTIDGEDGIC YTHPEKLPGV TRDGLSKHFF HRPSKAYTTG 120 TRKRRKIQTE CDVHKGETRW HKTGKTRPVM VNGRQKGCKK ILVLYTNFGK HRKPEKTNWV 180 MHQYHLGDLE EEKEGELVVC KIFYQTQPRQ CSWSSERGAA AAAAMAATTV QEQRRRDSGS 240 GSCSSRDHDV SATSFQAGYT VTTAVEMQQH MKQHSADHFS FAPFRKTFDQ EVGVGGDQVP 300 SNQLGRSEQH HAVQEQQPHR PVLATTAVPA TAFLISRPSN PVSTIVPPAM QHTSVVLDHD 360 QFHVPAILLH HHDKFQHQHQ QQQPPQKLDH RSAGLEELIM GCTSSTSTKG ETSIPQSQET 420 EWPYPYWPPD NQDHHG* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 3ulx_A | 3e-12 | 42 | 186 | 13 | 140 | Stress-induced transcription factor NAC1 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Pvr.21242 | 1e-175 | leaf| stem | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in the vascular cylinder of roots. Expressed in the differentiation zone of the root stele. {ECO:0000269|PubMed:25265867}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator involved in xylem formation (PubMed:25265867, PubMed:25535195). Promotes the expression of the secondary wall-associated transcription factor MYB46 (PubMed:25265867). Functions upstream of NAC030/VND7, a master switch of xylem vessel differentiation (PubMed:25265867, PubMed:25535195). Acts as upstream regulator of NAC101/VND6 and LBD30/ASL19 (PubMed:25535195). {ECO:0000269|PubMed:25265867, ECO:0000269|PubMed:25535195}. | |||||
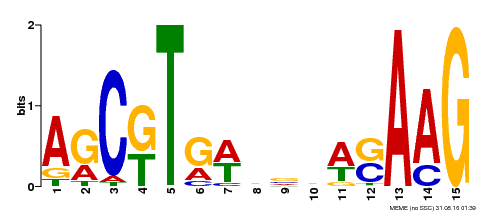
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00460 | DAP | Transfer from AT4G29230 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.5NG037700.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KM017005 | 0.0 | KM017005.1 UNVERIFIED: Miscanthus lutarioriparius NAC7-like mRNA, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025818126.1 | 0.0 | NAC domain-containing protein 75-like isoform X4 | ||||
| Swissprot | Q9M0F8 | 1e-149 | NAC75_ARATH; NAC domain-containing protein 75 | ||||
| TrEMBL | A0A2S3HYL8 | 0.0 | A0A2S3HYL8_9POAL; Uncharacterized protein | ||||
| TrEMBL | A0A2T7DS96 | 0.0 | A0A2T7DS96_9POAL; Uncharacterized protein | ||||
| STRING | Pavir.Eb00261.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1453 | 38 | 117 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G29230.1 | 1e-116 | NAC domain containing protein 75 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.5NG037700.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



