 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.5NG007800.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | Nin-like | ||||||||
| Protein Properties | Length: 936aa MW: 102671 Da PI: 6.1073 | ||||||||
| Description | Nin-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | RWP-RK | 91.8 | 5.2e-29 | 588 | 639 | 1 | 52 |
RWP-RK 1 aekeisledlskyFslpikdAAkeLgvclTvLKriCRqyGIkRWPhRkiksl 52
aek+isle+l++yFs+++k+AAk+Lgvc+T++KriCRq+GI+RWP+Rki+++
Pavir.5NG007800.1.p 588 AEKTISLEVLQQYFSGSLKNAAKSLGVCPTTMKRICRQHGISRWPSRKINKV 639
589***********************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51519 | 17.746 | 578 | 659 | IPR003035 | RWP-RK domain |
| Pfam | PF02042 | 3.3E-26 | 591 | 639 | IPR003035 | RWP-RK domain |
| SuperFamily | SSF54277 | 1.31E-20 | 831 | 926 | No hit | No description |
| SMART | SM00666 | 4.1E-24 | 841 | 923 | IPR000270 | PB1 domain |
| Pfam | PF00564 | 4.4E-16 | 841 | 922 | IPR000270 | PB1 domain |
| Gene3D | G3DSA:3.10.20.240 | 5.5E-25 | 841 | 922 | No hit | No description |
| PROSITE profile | PS51745 | 19.753 | 841 | 923 | IPR000270 | PB1 domain |
| CDD | cd06407 | 3.76E-32 | 842 | 922 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0010118 | Biological Process | stomatal movement | ||||
| GO:0010167 | Biological Process | response to nitrate | ||||
| GO:0042128 | Biological Process | nitrate assimilation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 936 aa Download sequence Send to blast |
MDLDPPTSGP GDACAVSADA WPFDSLTSSL LFPSVSASPP LPPLPANSSS WLTPPSPLWF 60 FEDRHLLPLD APQAPEATVA AAVVEEVQRA RSGNSDSTSK RVEQINHKWQ FHLSLDEDGT 120 DNSSLFKERL TQALRYFKDS TDQHLLVQVW APVKNGDRYV LTTSGQPFVL DHQSIGLLQY 180 RAVSMMYMFS VDGENVGELG LPGRVYKLKV PEWTPNVQYY SSTEFPRLNH AISYNVHGTV 240 ALPVFDPSAQ SCIAVVELIM TSKKINYACE VDKVCKALEA VNLKSTEILD HPNVQICNEG 300 RQTALVEILE ILTVVCEEHK LPLAQTWVPC KYRSVLAHGG GLKKSCLSFD GSCMGEVCMS 360 TSDVAFHVID AHMWGFRDAC VEHHLQRGQG VSGKAFITRK PCFSKDIRKF CKLKYPLVHY 420 ARMFGLAGCF AICLQSSYTG RDDYILEFFL PPDCIDEHDQ NALLESIFTL MKQCLRSLKV 480 VGDRDSSGVS LQLSNVLKIE NEEFKTDAQF DNSDGSLHES PKGDTHGGAH TFDNGDNNVP 540 NLPEGHLMAD DCSQDNGTSV GRPNGSGASD SLVLPKTSKP PERRRGKAEK TISLEVLQQY 600 FSGSLKNAAK SLGVCPTTMK RICRQHGISR WPSRKINKVN RSLSKLKQVI ESVQGSDAAF 660 NLTSITGPLP IPVGPSSDSL NVEKVTQNKV AEPSNLVVDG DRDSSLQKSL ENDGHFSILM 720 AQQGFIDNNN DSQLEADKAS RSRSSSGEGS INSRTSEGSC QGSPANRTFV CKPIASTFAE 780 PQLNPEEFHK EHFQEPQLPL SRMLIEDSGS SKDLKNLFTP AADQPFLAPP SNLVSMKHSG 840 TVTIKASFKE DIVRFRFPCS GSITVLKNEV AKRLRMDVGT FDIKYLDDDH EWVKLACNAD 900 LEECMEISRN SGSDVIRLLV SDITAHLGSS CGSSG* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Pvr.5084 | 0.0 | callus| leaf| stem | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
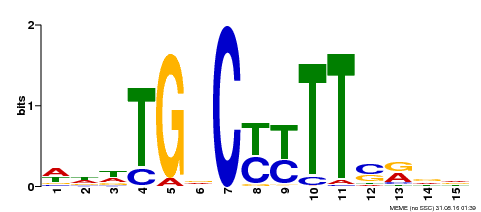
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00449 | DAP | Transfer from AT4G24020 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.5NG007800.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025814939.1 | 0.0 | protein NLP3 isoform X1 | ||||
| Swissprot | Q5NB82 | 0.0 | NLP3_ORYSJ; Protein NLP3 | ||||
| TrEMBL | A0A2T7DTG5 | 0.0 | A0A2T7DTG5_9POAL; Uncharacterized protein | ||||
| STRING | Si000216m | 0.0 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP4778 | 36 | 60 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G24020.1 | 0.0 | NIN like protein 7 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.5NG007800.1.p |



