 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.3KG460200.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 511aa MW: 54978.1 Da PI: 8.2641 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 47.2 | 5.1e-15 | 120 | 167 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g W++eEde+lv + + G g+W+ +ar g+ R++k+c++rw +yl
Pavir.3KG460200.1.p 120 KGLWSPEEDEKLVAYMLRSGQGSWSDVARNAGLQRCGKSCRLRWINYL 167
678*******************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 51.8 | 1.9e-16 | 173 | 216 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
rg+++++E+el+v +++ lG++ W+ Ia++++ gRt++++k++w++
Pavir.3KG460200.1.p 173 RGAFSPQEEELIVSLHAILGNR-WSQIAARLP-GRTDNEIKNFWNS 216
89********************.*********.***********97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 15.027 | 115 | 167 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 3.13E-28 | 118 | 214 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 8.7E-11 | 119 | 169 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.9E-13 | 120 | 167 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 4.0E-22 | 121 | 174 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 5.46E-9 | 123 | 167 | No hit | No description |
| PROSITE profile | PS51294 | 25.554 | 168 | 222 | IPR017930 | Myb domain |
| SMART | SM00717 | 9.6E-16 | 172 | 220 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 8.0E-15 | 173 | 216 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.96E-11 | 175 | 215 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 2.8E-25 | 175 | 222 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:1901348 | Biological Process | positive regulation of secondary cell wall biogenesis | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 511 aa Download sequence Send to blast |
MSSPSSFSSS FHSSSHYSMY TTNTLTHTHI TIYIQHTLRP ISSLSSTPTH TACSEIIDRP 60 HAHSLALLLD RRSYAHRRRP HVRVRDHHQG RTCRTAMRKP EGPAAANSCN NAGAAAKLRK 120 GLWSPEEDEK LVAYMLRSGQ GSWSDVARNA GLQRCGKSCR LRWINYLRPD LKRGAFSPQE 180 EELIVSLHAI LGNRWSQIAA RLPGRTDNEI KNFWNSTIKK RIKNSSSASS PAAATDCASP 240 EPNNGKVAGF DISGAASCPD LAGLDRQVDG GHHHAMVTTG LWMVDSSSST SSSTSPMQQG 300 RPPSSAMVAA VTRSYGGLLP LPDQLRGGMA ADASPAGFFH GHAAPFKHQA VVASLHGGYY 360 GGSAPHHHGM MATMEGGGGC FMRGEGLFGV PPLLEAMSSQ DQDHHGQALM ASSGGGNNNN 420 PKNNSSSNTT EATATTTTTV SNNESNITDN TKENINTMSL VNSSSSNVAA VYWEGAHQQY 480 MSRNVMHGEW DLEELMKDVS SLPFLDFQVE * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 2e-28 | 118 | 222 | 5 | 108 | B-MYB |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Pvr.18885 | 0.0 | stem | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
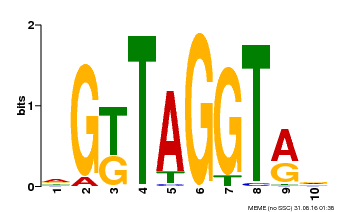
| Motif ID | Method | Source | Motif file |
| MP00599 | PBM | Transfer from AT5G12870 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.3KG460200.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KT075095 | 0.0 | KT075095.1 Panicum virgatum secondary wall-associated MYB domain-containing protein (MYB46B) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025808414.1 | 0.0 | myb-related protein 330-like | ||||
| TrEMBL | A0A0K1TPX6 | 0.0 | A0A0K1TPX6_PANVG; Secondary wall-associated MYB domain-containing protein | ||||
| STRING | Pavir.Ca02370.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP9821 | 35 | 45 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G12870.1 | 2e-64 | myb domain protein 46 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.3KG460200.1.p |



