 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.2NG401700.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 277aa MW: 29542.7 Da PI: 4.5123 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 60.1 | 3.4e-19 | 49 | 102 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k++++++eq+++Le+ Fe ++++ e++++LA+ lgL+ rqV vWFqNrRa++k
Pavir.2NG401700.1.p 49 KKRRLSAEQVRALERSFEVENKLEPERKARLARDLGLQPRQVAVWFQNRRARWK 102
45689************************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 133.7 | 6.9e-43 | 48 | 140 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLree 90
ekkrrls+eqv++LE+sFe e+kLeperK++lar+LglqprqvavWFqnrRAR+ktkqlE+dy+aL+++ydal++++++L++++++L +e
Pavir.2NG401700.1.p 48 EKKRRLSAEQVRALERSFEVENKLEPERKARLARDLGLQPRQVAVWFQNRRARWKTKQLERDYAALRHSYDALRRDHDALRRDKDALLAE 137
69**************************************************************************************99 PP
HD-ZIP_I/II 91 lke 93
+ke
Pavir.2NG401700.1.p 138 IKE 140
987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.03E-19 | 41 | 106 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.475 | 44 | 104 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.5E-18 | 47 | 108 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 1.6E-16 | 49 | 102 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 9.96E-19 | 49 | 105 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 3.0E-20 | 50 | 111 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 9.4E-6 | 75 | 84 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 79 | 102 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 9.4E-6 | 84 | 100 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 3.9E-17 | 104 | 145 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009637 | Biological Process | response to blue light | ||||
| GO:0030308 | Biological Process | negative regulation of cell growth | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048510 | Biological Process | regulation of timing of transition from vegetative to reproductive phase | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 277 aa Download sequence Send to blast |
MKRPRAGSPS LLPTASPDDD GYGAAAGMET EGDAEEEMVA CGGGGGGEKK RRLSAEQVRA 60 LERSFEVENK LEPERKARLA RDLGLQPRQV AVWFQNRRAR WKTKQLERDY AALRHSYDAL 120 RRDHDALRRD KDALLAEIKE LKAKLGDEEA AASFTSVKAE PAASDGPPAA GVGSSESDSS 180 AVLNDADPAV GAAEAPVPEV QGALLPAPTG AAATSHGGAF FHGGFLKVEE DETGFLDDDE 240 PCGGFFAVEQ PPPMAWWTEP TAGTELGWMR AQTWDL* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 73 | 79 | ERKARLA |
| 2 | 96 | 104 | RRARWKTKQ |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Pvr.1623 | 0.0 | callus| leaf| root| stem | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in leaf and floral organ primordia, floral meristems, embryonic axis and cells surrounding the vascular bundles. Expressed in the vasculature of roots, stem, leaves and spikelets, and in the vascular bundle of the scutellum in embryos. {ECO:0000269|PubMed:10732669, ECO:0000269|PubMed:17999151, ECO:0000269|PubMed:18049796}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in leaf and floral organ primordia, floral meristems, embryonic axis and cells surrounding the vascular bundles. Expressed in the vasculature of roots, stem, leaves and spikelets, and in the vascular bundle of the scutellum in embryos. {ECO:0000269|PubMed:10732669, ECO:0000269|PubMed:17999151, ECO:0000269|PubMed:18049796}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
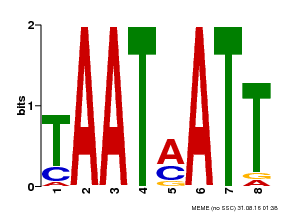
| UniProt | Probable transcription activator that binds to the DNA sequence 5'-CAAT[AT]ATTG-3'. May be involved in the regulation of gibberellin signaling. {ECO:0000269|PubMed:10732669, ECO:0000269|PubMed:18049796}. | |||||
| UniProt | Probable transcription activator that binds to the DNA sequence 5'-CAAT[AT]ATTG-3'. May be involved in the regulation of gibberellin signaling. {ECO:0000269|PubMed:10732669, ECO:0000269|PubMed:18049796}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00051 | PBM | Transfer from AT4G40060 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.2NG401700.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By gibberellin. Down-regulated in leaves by drought stress. {ECO:0000269|PubMed:17999151, ECO:0000269|PubMed:18049796}. | |||||
| UniProt | INDUCTION: By gibberellin. Down-regulated in leaves by drought stress. {ECO:0000269|PubMed:17999151, ECO:0000269|PubMed:18049796}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT034257 | 0.0 | BT034257.1 Zea mays full-length cDNA clone ZM_BFc0009H14 mRNA, complete cds. | |||
| GenBank | BT039062 | 0.0 | BT039062.1 Zea mays full-length cDNA clone ZM_BFb0367N11 mRNA, complete cds. | |||
| GenBank | BT055374 | 0.0 | BT055374.1 Zea mays full-length cDNA clone ZM_BFb0091J03 mRNA, complete cds. | |||
| GenBank | JX469929 | 0.0 | JX469929.1 Zea mays subsp. mays clone UT3187 HB-type transcription factor mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004957072.1 | 1e-162 | homeobox-leucine zipper protein HOX4 | ||||
| Swissprot | Q6K498 | 1e-123 | HOX4_ORYSJ; Homeobox-leucine zipper protein HOX4 | ||||
| Swissprot | Q9XH37 | 1e-123 | HOX4_ORYSI; Homeobox-leucine zipper protein HOX4 | ||||
| TrEMBL | K3ZW40 | 1e-160 | K3ZW40_SETIT; Uncharacterized protein | ||||
| STRING | Pavir.J27155.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3009 | 34 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G01470.1 | 2e-33 | homeobox 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.2NG401700.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



