 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.2KG306600.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 328aa MW: 34085 Da PI: 7.9016 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 55.7 | 1.2e-17 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT+eEd+ lv +v+++G+g+W++++ g+ R+ k+c++rw +yl
Pavir.2KG306600.1.p 14 KGPWTPEEDLMLVSYVQEHGPGNWRAVPTNTGLMRCSKSCRLRWTNYL 61
79********************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 50.7 | 4.3e-16 | 67 | 112 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg++T +E++l++++ ++lG++ W++Ia++++ Rt++++k++w+++l
Pavir.2KG306600.1.p 67 RGNFTDQEEKLIIHLQALLGNR-WAAIASYLP-ERTDNDIKNYWNTHL 112
89********************.*********.************996 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 3.1E-24 | 5 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 24.935 | 9 | 65 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.35E-30 | 11 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 7.5E-14 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.3E-16 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.85E-11 | 16 | 61 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 1.0E-24 | 65 | 117 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 20.592 | 66 | 116 | IPR017930 | Myb domain |
| SMART | SM00717 | 2.5E-16 | 66 | 114 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 4.2E-15 | 67 | 112 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.50E-11 | 69 | 112 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0080167 | Biological Process | response to karrikin | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 328 aa Download sequence Send to blast |
MGRPPCCDKV GVKKGPWTPE EDLMLVSYVQ EHGPGNWRAV PTNTGLMRCS KSCRLRWTNY 60 LRPGIKRGNF TDQEEKLIIH LQALLGNRWA AIASYLPERT DNDIKNYWNT HLKKKLKKMQ 120 AGEGGGGDDG AAGAASGSDG GDAAGDRTGG AGAGKRPAVP KGQWERRLQT DIHTARQALR 180 DALSLEPSVT PAKVAPPPTP PGSTTYASSA ENIARLLEGW LRPGGGKGPE ASGSTSTTAT 240 TQQRPQCSGE GAASASASHS GGAAGNTAAQ TPDYSTETSK MAGSAAGAGS APAFSMLESW 300 LLDDGMGHGE VGLMADVVPL GDPSEFF* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1mse_C | 7e-27 | 14 | 116 | 4 | 105 | C-Myb DNA-Binding Domain |
| 1msf_C | 7e-27 | 14 | 116 | 4 | 105 | C-Myb DNA-Binding Domain |
| Search in ModeBase | ||||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Expression in flowers increases as the flowers develop. {ECO:0000269|PubMed:1840903}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in flowers, leaves and weakly in seed pods. {ECO:0000269|PubMed:1840903}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor. {ECO:0000305}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00395 | DAP | Transfer from AT3G47600 | Download |
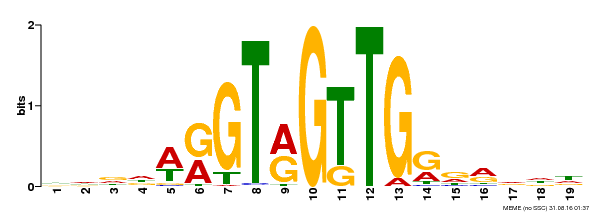
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.2KG306600.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HF679433 | 1e-165 | HF679433.1 Saccharum hybrid cultivar Co 86032 mRNA for ScMYB27 protein. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008652703.1 | 0.0 | myb-related protein 306 | ||||
| Swissprot | P81392 | 1e-109 | MYB06_ANTMA; Myb-related protein 306 | ||||
| TrEMBL | A0A096QJ49 | 0.0 | A0A096QJ49_MAIZE; Transcription factor MYB30 | ||||
| TrEMBL | A0A3L6DZ53 | 0.0 | A0A3L6DZ53_MAIZE; Myb-related protein 306 | ||||
| STRING | Pavir.Bb02186.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP229 | 38 | 296 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G47600.1 | 8e-90 | myb domain protein 94 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.2KG306600.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



