 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Potri.003G189700.4 | ||||||||
| Common Name | POPTR_0003s18900g | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Populus
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 559aa MW: 61063.7 Da PI: 4.7135 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 54.8 | 2.2e-17 | 39 | 86 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT Ed +l+++vk++G g+W+++ ++ g+ R++k+c++rw ++l
Potri.003G189700.4 39 KGPWTSAEDAILIEYVKKHGEGNWNSVQKHSGLFRCGKSCRLRWANHL 86
79******************************************9986 PP
| |||||||
| 2 | Myb_DNA-binding | 52.3 | 1.3e-16 | 92 | 135 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
+g++T eE++l++++++++G++ W++ a++++ gRt++++k++w++
Potri.003G189700.4 92 KGAFTHEEEQLIIELHAKMGNK-WARMAAHLP-GRTDNEIKNYWNT 135
799*******************.*********.***********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 17.172 | 34 | 86 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 8.61E-31 | 36 | 133 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.5E-15 | 38 | 88 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.5E-15 | 39 | 86 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 6.9E-24 | 40 | 93 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 4.17E-12 | 41 | 86 | No hit | No description |
| PROSITE profile | PS51294 | 26.568 | 87 | 141 | IPR017930 | Myb domain |
| SMART | SM00717 | 1.1E-16 | 91 | 139 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.1E-15 | 92 | 135 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 8.0E-26 | 94 | 140 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 3.71E-12 | 94 | 135 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009789 | Biological Process | positive regulation of abscisic acid-activated signaling pathway | ||||
| GO:0043068 | Biological Process | positive regulation of programmed cell death | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0045926 | Biological Process | negative regulation of growth | ||||
| GO:0048235 | Biological Process | pollen sperm cell differentiation | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 559 aa Download sequence Send to blast |
MSRTTSESED GMISKDQTGL PLGEEGSYGG STNGVGLKKG PWTSAEDAIL IEYVKKHGEG 60 NWNSVQKHSG LFRCGKSCRL RWANHLRPNL KKGAFTHEEE QLIIELHAKM GNKWARMAAH 120 LPGRTDNEIK NYWNTRIKRR QRAGLPLYPP EVSLQTLQGS QQCLDINGMD SGNKGQHDIL 180 QTHNYGIPDV MFDNLKTNRS ILPYVPELPD ISASSILMKG LCSSQYGSFM SPTMHRQKRL 240 REATTLLSSF SGGMKNDFHL FDQFQDGDKA AQYFGLSFPF DPDSATKNPE IFGENQGSHT 300 LANGDFSASK PTSEAVKFEL PSLQYAETDL GGWGASCSPS PLIESVDTFI QSPPTGTVES 360 NFPSPRNSGL LDALLYEART LSSAKNQSSD KSSNSSTITP GDNADCSALN ISETEWEDYG 420 DPISPLGHPA ASLFSECTPI SASGSSLDES PPTETLTECN MKLEPVDHTW TADREKESYT 480 QLDITRPDAL LDSDWLEHDS GYGKDQVIMT EAIATLLGDD LSSEYKQMAA GASTDHGWGL 540 GSCSWNNMPA VCQMSELP* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 8e-32 | 37 | 140 | 5 | 107 | B-MYB |
| Search in ModeBase | ||||||
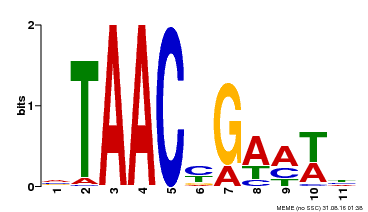
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00490 | DAP | Transfer from AT5G06100 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Potri.003G189700.4 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024452281.1 | 0.0 | transcription factor MYB33 isoform X2 | ||||
| TrEMBL | A0A3N7G7S8 | 0.0 | A0A3N7G7S8_POPTR; Uncharacterized protein | ||||
| STRING | POPTR_0003s18900.1 | 0.0 | (Populus trichocarpa) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G06100.3 | 1e-140 | myb domain protein 33 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Potri.003G189700.4 |
| Entrez Gene | 7461325 |



