 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Potri.001G347200.1 | ||||||||
| Common Name | POPTR_0001s35140g | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Populus
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 451aa MW: 50935.7 Da PI: 6.0138 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 53.2 | 6.9e-17 | 212 | 258 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g WT++Ed llv+ vkq+G + W+ Ia+++ gR +kqc++rw+++l
Potri.001G347200.1 212 KGQWTPQEDRLLVQSVKQYGIKKWSQIAKMLE-GRVGKQCRERWHNHL 258
799*****************************.*************97 PP
| |||||||
| 2 | Myb_DNA-binding | 57.7 | 2.8e-18 | 264 | 306 | 1 | 45 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwq 45
+++W++eEdell++a+++ G++ W+ Ia++++ gRt++ +k++w+
Potri.001G347200.1 264 KDAWSEEEDELLINAHREIGNR-WAEIAKRLP-GRTENTIKNHWN 306
679*******************.*********.***********8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 26.689 | 207 | 262 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.66E-31 | 210 | 305 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 9.3E-16 | 211 | 260 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 6.8E-28 | 213 | 265 | IPR009057 | Homeodomain-like |
| Pfam | PF13921 | 1.0E-18 | 215 | 272 | No hit | No description |
| CDD | cd00167 | 7.72E-15 | 215 | 258 | No hit | No description |
| PROSITE profile | PS51294 | 21.306 | 263 | 313 | IPR017930 | Myb domain |
| SMART | SM00717 | 4.4E-17 | 263 | 311 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.4E-22 | 266 | 311 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 2.10E-13 | 266 | 309 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010439 | Biological Process | regulation of glucosinolate biosynthetic process | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:1904095 | Biological Process | negative regulation of endosperm development | ||||
| GO:2000692 | Biological Process | negative regulation of seed maturation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 451 aa Download sequence Send to blast |
MELNTMLIDE FPFLSSLLSE YNLNPEGTND FSLEEALSSK GLFYNFHHLG YGHPTNIIPP 60 YLDLDHFTIE GSSQNPFSGI SGTCIDPARL DSLASGFSSH DLNAYTPTVV PLLPAGCGDR 120 LLHGLQRGSV RDHDYQKISG ARSLNQKEMK EQRGFEEIGR TRTANNGVST DEASCVSTED 180 SKHHKQVDHH RKAKKLLLEK DSKVHKKSQV IKGQWTPQED RLLVQSVKQY GIKKWSQIAK 240 MLEGRVGKQC RERWHNHLRP DIKKDAWSEE EDELLINAHR EIGNRWAEIA KRLPGRTENT 300 IKNHWNATKR RQFSRRESGK DLDSKSTLLQ SYIMMVTSSS STQENNEEEK TDDVNSNSPN 360 QNESPGSSST DLEIPYGHGH NNEASKLSFD TNLFNDSYGF MSFLEEMPCS CVVDESNMEF 420 EISGLDSLMK GAEVKKEMDL LEMITQGISH * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1mse_C | 1e-44 | 210 | 312 | 2 | 104 | C-Myb DNA-Binding Domain |
| 1msf_C | 1e-44 | 210 | 312 | 2 | 104 | C-Myb DNA-Binding Domain |
| Search in ModeBase | ||||||
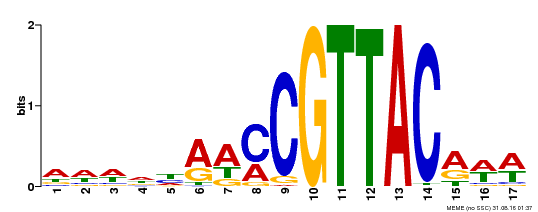
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00385 | DAP | Transfer from AT3G27785 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Potri.001G347200.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002300044.1 | 0.0 | transcription factor MYB118 | ||||
| TrEMBL | B9GI43 | 0.0 | B9GI43_POPTR; Uncharacterized protein | ||||
| STRING | POPTR_0001s35140.1 | 0.0 | (Populus trichocarpa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1570 | 31 | 96 | Representative plant | OGRP5 | 17 | 1784 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G27785.1 | 6e-70 | myb domain protein 118 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Potri.001G347200.1 |
| Entrez Gene | 7496834 |



