 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Potri.001G188800.2 | ||||||||
| Common Name | HB5, POPTR_0001s18930g | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Populus
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 840aa MW: 92429.7 Da PI: 6.4251 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 57.7 | 2.1e-18 | 19 | 77 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+l++ +++ps +r++L +++ +++ +q+kvWFqNrR +ek+
Potri.001G188800.2 19 KYVRYTPEQVEALERLYHDCPKPSSIRRQQLIRECpilsNIEPKQIKVWFQNRRCREKQ 77
5679*****************************************************97 PP
| |||||||
| 2 | START | 175.8 | 2.7e-55 | 163 | 371 | 2 | 205 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEEECTT..EE CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetlevissg..ga 90
+aee+++e+++ka+ ++ Wv+++ +++g++++ +++ s+++sg +ra+g+v +++ v+e+l+d++ W ++++++++l+v+ ++ g+
Potri.001G188800.2 163 IAEETLTEFLSKATGTAVEWVQMPGMKPGPDSIGIVAISHGCSGVGARACGLVGLEPT-RVAEILKDRPSWFRDCRAVDVLNVLPTAngGT 252
789*******************************************************.8888888888****************9999** PP
EEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXX CS
START 91 lqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvtwvehvdlkgrlp 177
++l +++l+a+++l+p Rdf+ +Ry+ l++g++v++++S+ ++q+ p+ +++vRae+lpSg+l++p+++g+s +++v+h+dl+ +++
Potri.001G188800.2 253 IELLYMQLYAPTTLAPgRDFWLLRYTSVLEDGSLVVCERSLKNTQNGPSmppVQHFVRAEMLPSGYLVRPCEGGGSIIHIVDHMDLEPWSV 343
***********************************************9988899************************************* PP
HHHHHHHHHHHHHHHHHHHHHHTXXXXX CS
START 178 hwllrslvksglaegaktwvatlqrqce 205
+++lr+l++s+++ ++kt++a+l+++++
Potri.001G188800.2 344 PEVLRPLYESSTVLAQKTTMAALRQLRQ 371
************************9876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 15.613 | 14 | 78 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.2E-15 | 16 | 82 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 1.33E-16 | 18 | 82 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 1.32E-16 | 19 | 79 | No hit | No description |
| Pfam | PF00046 | 5.8E-16 | 20 | 77 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 3.5E-18 | 21 | 77 | IPR009057 | Homeodomain-like |
| CDD | cd14686 | 1.27E-6 | 71 | 110 | No hit | No description |
| PROSITE profile | PS50848 | 24.652 | 153 | 368 | IPR002913 | START domain |
| CDD | cd08875 | 4.39E-77 | 157 | 373 | No hit | No description |
| Gene3D | G3DSA:3.30.530.20 | 1.4E-22 | 162 | 369 | IPR023393 | START-like domain |
| SuperFamily | SSF55961 | 2.88E-37 | 162 | 374 | No hit | No description |
| SMART | SM00234 | 5.5E-39 | 162 | 372 | IPR002913 | START domain |
| Pfam | PF01852 | 9.0E-53 | 163 | 371 | IPR002913 | START domain |
| Pfam | PF08670 | 1.3E-52 | 696 | 838 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 840 aa Download sequence Send to blast |
MMAMSCKDGK NPINMDNGKY VRYTPEQVEA LERLYHDCPK PSSIRRQQLI RECPILSNIE 60 PKQIKVWFQN RRCREKQRKE ASRLQAVNRK LTAMNKLLME ENDRLQKQVS QLVYENGYFR 120 QHTQNTTLAS KDTSCESVVT SGQHHLTPQH PPRDASPAGL LSIAEETLTE FLSKATGTAV 180 EWVQMPGMKP GPDSIGIVAI SHGCSGVGAR ACGLVGLEPT RVAEILKDRP SWFRDCRAVD 240 VLNVLPTANG GTIELLYMQL YAPTTLAPGR DFWLLRYTSV LEDGSLVVCE RSLKNTQNGP 300 SMPPVQHFVR AEMLPSGYLV RPCEGGGSII HIVDHMDLEP WSVPEVLRPL YESSTVLAQK 360 TTMAALRQLR QIAQEASQSS VTNWGRRPAA LRALSQRLSR GFNEALNGFS DEGWSMIGND 420 GMDDVTILVN SSPDKLMGLN LSFSNGFPAV SSAVLCAKAS MLLQNVPPAI LLRFLREHRS 480 EWADNNIDAY AAAAVKVGPC SLQGSRVGNF GGQVILPLAH TVEHEEFLEV IKLEGVCHSP 540 EDAIMPRDVF LLQLCCGMDE NAVGTCAELI FAPIDATFAD DAPLLPSGFR IIPLDSGKEA 600 SSPNRTLDLA SALEVGAGNR ASSDFSANSG CTRSVMTIAF EFAFESHMQE HVASMARQYI 660 RSIISSVQRV ALALSPSHQG SQAGLRSPLG TPEAQTLARW ICQSYRNYLG VELLKSSSEG 720 SESILKTLWH HSDAIMCCSL KALPVFTFAN QAGLDMLETT LVALQDITLE KIFDDHGRKT 780 LCSEFPQIMQ QGFTCLQGGI CLSSMGRPVS YERAVSWKVL NEEENAHCIC FMFINWSFV* |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Highly expressed the developing vascular elements and the adaxial portion of cotyledons. Expressed in developing ovules, stamens and carpels. Expressed in procambium and shoot meristem. {ECO:0000269|PubMed:14701930, ECO:0000269|PubMed:15705957}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of meristem development to promote lateral organ formation. May regulates procambial and vascular tissue formation or maintenance, and vascular development in inflorescence stems. {ECO:0000269|PubMed:15598805, ECO:0000269|PubMed:15705957, ECO:0000269|PubMed:16617092}. | |||||
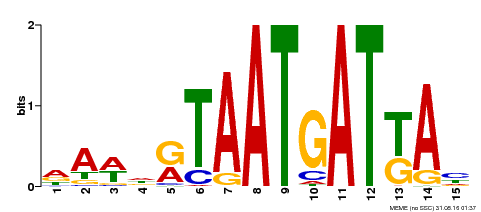
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Potri.001G188800.2 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. Repressed by miR165 and miR166. {ECO:0000269|PubMed:15773855, ECO:0000269|PubMed:16033795, ECO:0000269|PubMed:16617092, ECO:0000269|PubMed:17237362}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KF534716 | 0.0 | KF534716.1 Populus alba x Populus glandulosa HD-ZIPIII mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024464085.1 | 0.0 | homeobox-leucine zipper protein ATHB-15 | ||||
| Swissprot | Q9ZU11 | 0.0 | ATB15_ARATH; Homeobox-leucine zipper protein ATHB-15 | ||||
| TrEMBL | A0A2I4KKW3 | 0.0 | A0A2I4KKW3_POPTR; HD-ZIP family protein | ||||
| STRING | POPTR_0001s18930.1 | 0.0 | (Populus trichocarpa) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.1 | 0.0 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Potri.001G188800.2 |
| Entrez Gene | 7470577 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



