 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | model.Picochlorum_contig_87.g285.t1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; Trebouxiophyceae; Trebouxiophyceae incertae sedis; Picochlorum
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 504aa MW: 54648 Da PI: 6.307 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 87 | 1.9e-27 | 187 | 240 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kpr++W+ e+H++Fv av+ L G++kA+Pk+il+lm+v+gLt+e+v+SHLQkYRl
model.Picochlorum_contig_87.g285.t1 187 KPRVVWSVEMHQQFVYAVNTL-GIDKAVPKRILDLMNVEGLTRENVASHLQKYRL 240
79*******************.********************************8 PP
| |||||||
| 2 | Response_reg | 84.9 | 2.4e-28 | 20 | 131 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH.....ESEEEEESSCTTSEHHHHHHHH.HHH CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd.....pDlillDiempgmdGlellkei.ree 68
vl+vdD+p ++++ +l++ +y ev++ ++g++al++l++++ +Dl+l D+ mp+mdG++ll++i e
model.Picochlorum_contig_87.g285.t1 20 VLVVDDDPMCLKVVSAMLKRCNY-EVDTESNGHQALDKLRQMQengcqFDLVLSDVYMPDMDGFKLLESIgLEL 92
89*********************.***************999999*************************5554 PP
TTTSEEEEEESTTTHHHHHHHHHTTESEEEESS--HHHHHH CS
Response_reg 69 epklpiivvtahgeeedalealkaGakdflsKpfdpeelvk 109
++p+i++++ ++ + +l+ + Ga dfl Kp+ +eel++
model.Picochlorum_contig_87.g285.t1 93 --EVPVIMMSSDSDTNVVLRGVTHGAVDFLIKPVRIEELRN 131
..6************************************87 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF036392 | 2.6E-121 | 1 | 475 | IPR017053 | Response regulator B-type, plant |
| SuperFamily | SSF52172 | 3.56E-37 | 17 | 145 | IPR011006 | CheY-like superfamily |
| Gene3D | G3DSA:3.40.50.2300 | 5.5E-41 | 17 | 153 | No hit | No description |
| SMART | SM00448 | 1.0E-31 | 18 | 133 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 43.684 | 19 | 137 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 6.1E-25 | 20 | 131 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 9.87E-31 | 21 | 137 | No hit | No description |
| SuperFamily | SSF46689 | 3.94E-19 | 185 | 244 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 1.8E-28 | 186 | 245 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.0E-23 | 187 | 240 | IPR006447 | Myb domain, plants |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009736 | Biological Process | cytokinin-activated signaling pathway | ||||
| GO:0009873 | Biological Process | ethylene-activated signaling pathway | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0080113 | Biological Process | regulation of seed growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 504 aa Download sequence Send to blast |
MSRQEAGGSE GTFSPAGLKV LVVDDDPMCL KVVSAMLKRC NYEVDTESNG HQALDKLRQM 60 QENGCQFDLV LSDVYMPDMD GFKLLESIGL ELEVPVIMMS SDSDTNVVLR GVTHGAVDFL 120 IKPVRIEELR NVWQHVIRRR SSQMSRGATA EESGVEQDIN SRTSQGTKRK DVEVIRAEHE 180 GGSTGKKPRV VWSVEMHQQF VYAVNTLGID KAVPKRILDL MNVEGLTREN VASHLQKYRL 240 YLKRVEGVAG KGNVAVVKGG VPRNVDRLNG KPPLQNKVPK QSKSCGTSRA ESSDAERKLN 300 TLHASGSSPN LQGISSSHWS QPHGYYNLHG QIYGNSVRMN ATGAPPTMMG WQNQNGVPMA 360 TPIQYANVPS TSAFYQGSGH MYPAQVPPIN GQMGVTNNNN VLHSAIDSSQ RPSCTEAHVN 420 GSTNATGNAP GSDNEFDTVV NNNTNVHSST DDASHNKIAT DILRSGSSGL DDDGLPNIAG 480 LDKNHDPDDD VLDMFIKGME GET* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 4e-22 | 186 | 245 | 4 | 63 | ARR10-B |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00054 | PBM | Transfer from AT4G16110 | Download |
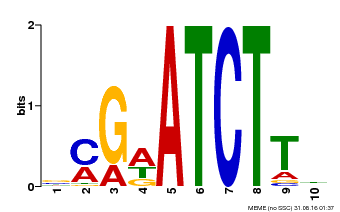
| |||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP1220 | 15 | 16 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G16110.1 | 3e-86 | response regulator 2 | ||||



