 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Prupe.7G183300.2.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Amygdaleae; Prunus
|
||||||||
| Family | WOX | ||||||||
| Protein Properties | Length: 255aa MW: 29269.8 Da PI: 6.405 | ||||||||
| Description | WOX family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 69.4 | 4.4e-22 | 89 | 149 | 2 | 57 |
T--SS--HHHHHHHHHHHHH.SSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 2 rkRttftkeqleeLeelFek.nrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
r+R+t+t+ ql++Le+lFe+ n +ps+++++e++++l +++e++V++WFqNrRa+ k+
Prupe.7G183300.2.p 89 RQRWTPTPVQLQILERLFEQgNGTPSKQKIKEITSELsqhgQISETNVYNWFQNRRARSKR 149
89*****************99**************************************97 PP
| |||||||
| 2 | Wus_type_Homeobox | 101.8 | 4.8e-33 | 88 | 151 | 2 | 65 |
Wus_type_Homeobox 2 artRWtPtpeQikiLeelyksGlrtPnkeeiqritaeLeeyGkiedkNVfyWFQNrkaRerqkq 65
ar+RWtPtp Q++iLe+l+++G tP+k++i++it+eL+++G+i+++NV++WFQNr+aR+++kq
Prupe.7G183300.2.p 88 ARQRWTPTPVQLQILERLFEQGNGTPSKQKIKEITSELSQHGQISETNVYNWFQNRRARSKRKQ 151
789***********************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 1.5E-18 | 84 | 154 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 15.37 | 85 | 150 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 5.8E-15 | 87 | 154 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 2.69E-19 | 88 | 154 | IPR009057 | Homeodomain-like |
| Pfam | PF00046 | 7.2E-20 | 89 | 149 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 8.45E-13 | 89 | 151 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 255 aa Download sequence Send to blast |
MMEWEKLQQK EHEMDGQANG NGGVLYVKVM TDEQLETLRK QIAVYASICE QLVEMHKNLT 60 AQQDLAGVRM GNLYCDPLMT SAGHKITARQ RWTPTPVQLQ ILERLFEQGN GTPSKQKIKE 120 ITSELSQHGQ ISETNVYNWF QNRRARSKRK QQNAACNHAE SEVETEVDSP KDKKTRPEEL 180 FQSQQNSAQR TEDLCFQSPE ISSDLHFLDP QTTKTDKMFP SNSGLRTSRH LSEMSFYDEV 240 LSNSTEPQKF KLRE* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 142 | 150 | RRARSKRKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor which may be involved in developmental processes. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00617 | PBM | Transfer from AT4G35550 | Download |
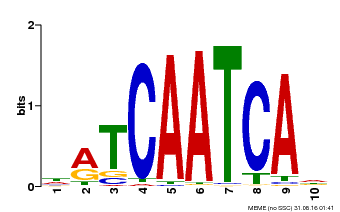
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Prupe.7G183300.2.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007203410.2 | 0.0 | WUSCHEL-related homeobox 8 | ||||
| Swissprot | O81788 | 6e-91 | WOX13_ARATH; WUSCHEL-related homeobox 13 | ||||
| TrEMBL | A0A251NDA4 | 0.0 | A0A251NDA4_PRUPE; Uncharacterized protein | ||||
| STRING | XP_008241933.1 | 1e-177 | (Prunus mume) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G35550.1 | 2e-93 | WUSCHEL related homeobox 13 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Prupe.7G183300.2.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



