 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Prupe.6G242700.2.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Amygdaleae; Prunus
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 284aa MW: 31417.4 Da PI: 9.3021 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 43.6 | 6.7e-14 | 36 | 80 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT eE++++++a +++ + Wk+I +g +t q++s+ qky
Prupe.6G242700.2.p 36 RESWTDEEHDKFLEALQLFDRD-WKKIEDFVG-SKTVIQIRSHAQKY 80
789*****************77.*********.*************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.2E-17 | 30 | 86 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 15.897 | 31 | 85 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.1E-7 | 34 | 86 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.4E-18 | 34 | 83 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 2.6E-11 | 35 | 83 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.2E-11 | 36 | 80 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 5.40E-9 | 38 | 81 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0032922 | Biological Process | circadian regulation of gene expression | ||||
| GO:0043966 | Biological Process | histone H3 acetylation | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 284 aa Download sequence Send to blast |
MNSTSSNNPQ SMGAASTPSD GSGKKVRKPY TITKSRESWT DEEHDKFLEA LQLFDRDWKK 60 IEDFVGSKTV IQIRSHAQKY FLKVQKSGTT AHVPPPRPKR KAAHPYPQKA SKNALVPLRA 120 SIPYPSSMNA IASTYSPWDE TSIMINPASN QILLSHEEFT SFQGTEADIR SKGLARINKC 180 SLSGIESSSR TLPSSEIPEQ GKQAPLLHGI PDFSEVYSFI GSVFDPDSKG HAQKLREMDP 240 INFETVLLLM RNLSINLSSP DFEPIVLFWS ENSGSSSRTA CRI* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator of evening element (EE)-containing clock-controlled genes. Forms a negative feedback loop with APRR5. Regulates the pattern of histone H3 acetylation of the TOC1 promoter. {ECO:0000269|PubMed:21205033, ECO:0000269|PubMed:21474993, ECO:0000269|PubMed:21483796, ECO:0000269|PubMed:23638299}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00338 | DAP | Transfer from AT3G09600 | Download |
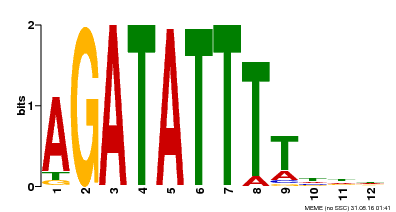
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Prupe.6G242700.2.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Circadian-regulation. Peak of expression in the afternoon. Down-regulated by cold. {ECO:0000269|PubMed:21205033, ECO:0000269|PubMed:22902701, ECO:0000269|PubMed:23638299}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB627311 | 1e-156 | AB627311.1 Malus x domestica mRNA, microsatellite: MEST187, clone: SAM01_15_D05. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020423139.1 | 0.0 | protein REVEILLE 8 | ||||
| Swissprot | Q8RWU3 | 1e-125 | RVE8_ARATH; Protein REVEILLE 8 | ||||
| TrEMBL | A0A251NV39 | 0.0 | A0A251NV39_PRUPE; Uncharacterized protein | ||||
| STRING | XP_008242131.1 | 0.0 | (Prunus mume) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G09600.1 | 1e-113 | MYB_related family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Prupe.6G242700.2.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



