 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Prupe.6G129100.6.p | ||||||||
| Common Name | PRUPE_ppa004537mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Amygdaleae; Prunus
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 516aa MW: 56915.7 Da PI: 6.6783 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 31.6 | 3.7e-10 | 212 | 252 | 5 | 45 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaL 45
k rr+++NReAAr+sR+RKka++++Le+ +L++ ++L
Prupe.6G129100.6.p 212 KTLRRLAQNREAARKSRLRKKAYVQQLETSRIKLTQLEQDL 252
788*************************9877777766666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 2.0E-7 | 208 | 271 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 9.174 | 210 | 254 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 1.37E-6 | 212 | 254 | No hit | No description |
| Pfam | PF00170 | 2.0E-7 | 212 | 253 | IPR004827 | Basic-leucine zipper domain |
| Gene3D | G3DSA:1.20.5.170 | 3.5E-7 | 213 | 255 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 215 | 230 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF14144 | 5.6E-32 | 293 | 368 | IPR025422 | Transcription factor TGA like domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 516 aa Download sequence Send to blast |
MAFISATCPD SSFFLEYKDE NGLRKSCMAS HRVGETATGL SESGPSNHHV PYAVLHGMNA 60 PSTSFLNQEG SAFDFGELEE AIARQVRNDE AQAPLFTGRP AATLEMFPSW PMRFHQTPRG 120 SSKSGGESTD SGSQVNTLTS KGEGGQLEPE SPISKKASSS DHQQTFDQKH LQFQQQQQLQ 180 AQDMAISDSS RGGAVGVGAS QSQSAAKPSQ EKTLRRLAQN REAARKSRLR KKAYVQQLET 240 SRIKLTQLEQ DLQRARAQGL FLGGCGGGLG NISSGAAIFD MEYARWLEDD HRHMSELRTG 300 LQAHLSDSDL RVIVDGYISH YDEIFQLKGV AAKSDVFHLI TGMWTTPAER CFLWMGGFRP 360 SELIKMLTAQ LDPLTEQQFM GIYSLQQSSQ QAEEALTQGL EQLHQSLVDT IAGGPVIDGM 420 QQMAVALGKL TNLEGFVRQA DNLRQQTLHQ LRRILTVRQA ARCFLVIGEY YGRLRALSSL 480 WASRPRESMI SDDNSCQTTT DLQMVQPSQN HFSSF* |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: During anther development, accumulates in anther primordia during archesporial cell specification and later present in a horseshoe pattern associated with the lateral and adaxial portion of primordia, prior to the emergence of distinct locules. Expressed throughout sporogenic tissue and surrounding cells layers in adaxial and adaxial locules. Localized to the tapetum and middle layers, gradually fading postmeiosis with degeneration of these cell layers. {ECO:0000269|PubMed:20805327}. | |||||
| Uniprot | TISSUE SPECIFICITY: Mostly expressed in stems, inflorescence apex and flowers, and, to a lower extent, in seedlings, leaves and siliques. {ECO:0000269|PubMed:20805327}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Together with TGA10, basic leucine-zipper transcription factor required for anther development, probably via the activation of SPL expression in anthers and via the regulation of genes with functions in early and middle tapetal development (PubMed:20805327). Required for signaling responses to pathogen-associated molecular patterns (PAMPs) such as flg22 that involves chloroplastic reactive oxygen species (ROS) production and subsequent expression of H(2)O(2)-responsive genes (PubMed:27717447). {ECO:0000269|PubMed:20805327, ECO:0000269|PubMed:27717447}. | |||||
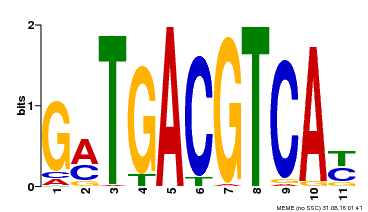
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00131 | DAP | Transfer from AT1G08320 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Prupe.6G129100.6.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by flg22 in leaves. {ECO:0000269|PubMed:27717447}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008228573.1 | 0.0 | PREDICTED: transcription factor TGA2 isoform X2 | ||||
| Swissprot | Q93XM6 | 0.0 | TGA9_ARATH; Transcription factor TGA9 | ||||
| TrEMBL | A0A251NPJ0 | 0.0 | A0A251NPJ0_PRUPE; Uncharacterized protein | ||||
| STRING | XP_008228563.1 | 0.0 | (Prunus mume) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G08320.3 | 0.0 | bZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Prupe.6G129100.6.p |
| Entrez Gene | 18774208 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



