 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Prupe.3G170600.1.p | ||||||||
| Common Name | MADS8, PRUPE_ppa010578mg, SHP | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Amygdaleae; Prunus
|
||||||||
| Family | MIKC_MADS | ||||||||
| Protein Properties | Length: 245aa MW: 28313.8 Da PI: 9.9887 | ||||||||
| Description | MIKC_MADS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SRF-TF | 98.5 | 2.6e-31 | 24 | 73 | 1 | 50 |
S---SHHHHHHHHHHHHHHHHHHHHHHHHHHT-EEEEEEE-TTSEEEEEE CS
SRF-TF 1 krienksnrqvtfskRrngilKKAeELSvLCdaevaviifsstgklyeys 50
krien++nrqvtf+kRrng+lKKA+ELSvLCdaeva+i+fs++g+lyey+
Prupe.3G170600.1.p 24 KRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSTRGRLYEYA 73
79***********************************************8 PP
| |||||||
| 2 | K-box | 112.8 | 3.4e-37 | 92 | 188 | 4 | 100 |
K-box 4 ssgksleeakaeslqqelakLkkeienLqreqRhllGedLesLslkeLqqLeqqLekslkkiRskKnellleqieelqkkekelqeenkaL 94
++g s++ea+++ +qqe++kL+++i+++q+++Rh+lGe L++L++keL++Le +Lek++++iRskKne+l+++ie +qk+e elq++n++L
Prupe.3G170600.1.p 92 TNGGSVSEANTQFYQQESSKLRRQIREIQNSNRHILGEALSTLNIKELKNLEGRLEKGISRIRSKKNEMLFAEIEFMQKREMELQNHNNYL 182
556669************************************************************************************* PP
K-box 95 rkklee 100
r+k++e
Prupe.3G170600.1.p 183 RAKIAE 188
***986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00432 | 2.0E-40 | 16 | 75 | IPR002100 | Transcription factor, MADS-box |
| PROSITE profile | PS50066 | 33.515 | 16 | 76 | IPR002100 | Transcription factor, MADS-box |
| SuperFamily | SSF55455 | 8.11E-33 | 17 | 89 | IPR002100 | Transcription factor, MADS-box |
| CDD | cd00265 | 5.64E-43 | 17 | 91 | No hit | No description |
| PROSITE pattern | PS00350 | 0 | 18 | 72 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 8.1E-33 | 18 | 38 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF00319 | 2.0E-26 | 25 | 72 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 8.1E-33 | 38 | 53 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 8.1E-33 | 53 | 74 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF01486 | 2.0E-26 | 100 | 186 | IPR002487 | Transcription factor, K-box |
| PROSITE profile | PS51297 | 15.203 | 102 | 192 | IPR002487 | Transcription factor, K-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 245 aa Download sequence Send to blast |
MEFPNQAPES SSQRKIGRGK IEIKRIENTT NRQVTFCKRR NGLLKKAYEL SVLCDAEVAL 60 IVFSTRGRLY EYANNSVRAT IDRYKKACTD STNGGSVSEA NTQFYQQESS KLRRQIREIQ 120 NSNRHILGEA LSTLNIKELK NLEGRLEKGI SRIRSKKNEM LFAEIEFMQK REMELQNHNN 180 YLRAKIAENE RAQQQQTNMI QGTSYDQSMP SQSYDRNFLP VILEANNNNN NHYSRHDQTA 240 LQLV* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1egw_A | 8e-20 | 17 | 84 | 1 | 68 | MADS BOX TRANSCRIPTION ENHANCER FACTOR 2, POLYPEPTIDE A |
| 1egw_B | 8e-20 | 17 | 84 | 1 | 68 | MADS BOX TRANSCRIPTION ENHANCER FACTOR 2, POLYPEPTIDE A |
| 1egw_C | 8e-20 | 17 | 84 | 1 | 68 | MADS BOX TRANSCRIPTION ENHANCER FACTOR 2, POLYPEPTIDE A |
| 1egw_D | 8e-20 | 17 | 84 | 1 | 68 | MADS BOX TRANSCRIPTION ENHANCER FACTOR 2, POLYPEPTIDE A |
| 1tqe_P | 9e-20 | 16 | 84 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| 1tqe_Q | 9e-20 | 16 | 84 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| 1tqe_R | 9e-20 | 16 | 84 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| 1tqe_S | 9e-20 | 16 | 84 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| 5f28_A | 8e-20 | 16 | 84 | 1 | 69 | MEF2C |
| 5f28_B | 8e-20 | 16 | 84 | 1 | 69 | MEF2C |
| 5f28_C | 8e-20 | 16 | 84 | 1 | 69 | MEF2C |
| 5f28_D | 8e-20 | 16 | 84 | 1 | 69 | MEF2C |
| 6byy_C | 9e-20 | 16 | 84 | 1 | 69 | MEF2 CHIMERA |
| 6byy_D | 9e-20 | 16 | 84 | 1 | 69 | MEF2 CHIMERA |
| 6c9l_A | 9e-20 | 16 | 84 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| 6c9l_B | 9e-20 | 16 | 84 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| 6c9l_C | 9e-20 | 16 | 84 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| 6c9l_D | 9e-20 | 16 | 84 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| 6c9l_E | 9e-20 | 16 | 84 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| 6c9l_F | 9e-20 | 16 | 84 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Ppe.2689 | 0.0 | fruit | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in flowers and seeds. {ECO:0000269|PubMed:11414613}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in flower development. {ECO:0000250|UniProtKB:Q0HA25}. | |||||
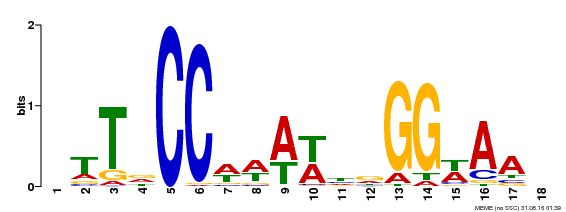
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00093 | SELEX | Transfer from AT3G58780 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Prupe.3G170600.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | DQ777635 | 0.0 | DQ777635.1 Prunus persica SHATTERPROOF-like (SHP) mRNA, complete cds. | |||
| GenBank | EU072119 | 0.0 | EU072119.1 Prunus persica SHATTERPROOF-like protein (MADS8) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007217264.1 | 0.0 | agamous-like MADS-box protein AGL1 | ||||
| Refseq | XP_020414870.1 | 0.0 | agamous-like MADS-box protein AGL1 | ||||
| Swissprot | Q93XH4 | 1e-128 | MADS1_VITVI; Agamous-like MADS-box protein MADS1 | ||||
| TrEMBL | Q0GPY8 | 1e-179 | Q0GPY8_PRUPE; PLENA-like MADS-box protein | ||||
| STRING | EMJ18463 | 1e-180 | (Prunus persica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF684 | 29 | 102 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G58780.1 | 1e-111 | MIKC_MADS family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Prupe.3G170600.1.p |
| Entrez Gene | 18783633 |



