 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Prupe.2G156900.2.p | ||||||||
| Common Name | PRUPE_ppa001436mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Amygdaleae; Prunus
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 796aa MW: 86560 Da PI: 6.2698 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 65.4 | 7.6e-21 | 124 | 179 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r eL+++l L++rqVk+WFqNrR+++k
Prupe.2G156900.2.p 124 KKRYHRHTPQQIQELEALFKECPHPDEKQRLELSRRLCLETRQVKFWFQNRRTQMK 179
688999***********************************************999 PP
| |||||||
| 2 | START | 30.5 | 7.9e-11 | 331 | 408 | 1 | 69 |
HHHHHHHHHHHHHHHHC-TT-EEEE....EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHHHHHHHHCCCG CS
START 1 elaeeaaqelvkkalaeepgWvkss....esengdevlqkfeeskv.....dsgealrasgvvdmvlallveellddk 69
ela++a++elvk+a+ +ep+W +s e +n +e++++f++ + + +ea+r++g+v+ ++ lve+l++++
Prupe.2G156900.2.p 331 ELALAAMDELVKLAQTDEPLWLRSLeggrEVLNHEEYMRSFTPCIGlkpngFVTEASRETGMVIINSLALVETLMESM 408
5899************************99***********9999899999************************987 PP
| |||||||
| 3 | START | 136.1 | 3.9e-43 | 408 | 521 | 94 | 206 |
EEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--.-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHHHHHH CS
START 94 mvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppesssvvRaellpSgiliepksnghskvtwvehvdlkgrlphwllrs 183
m aelq+lsplvp R++ f+R+++q+ +g+w++vdvSvd ++ + + ++ +++lpSg+++++++ng+skvtwveh++++++++h+l+r+
Prupe.2G156900.2.p 408 MHAELQVLSPLVPvREVNFLRFCKQHAEGVWAVVDVSVDTIRDTSGAPTFMNCRRLPSGCVVQDMPNGYSKVTWVEHAEYDESQVHQLYRP 498
789****************************************999********************************************* PP
HHHHHHHHHHHHHHHHTXXXXXX CS
START 184 lvksglaegaktwvatlqrqcek 206
+++sg+ +ga++wvatlqrqce+
Prupe.2G156900.2.p 499 MLSSGMGFGAQRWVATLQRQCEC 521
*********************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 3.98E-21 | 104 | 181 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.5E-22 | 108 | 181 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.524 | 121 | 181 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 5.7E-17 | 122 | 185 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 2.09E-18 | 124 | 181 | No hit | No description |
| Pfam | PF00046 | 2.1E-18 | 124 | 179 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 156 | 179 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 30.974 | 322 | 524 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.26E-26 | 323 | 521 | No hit | No description |
| CDD | cd08875 | 9.08E-99 | 326 | 520 | No hit | No description |
| SMART | SM00234 | 1.0E-24 | 331 | 521 | IPR002913 | START domain |
| Pfam | PF01852 | 1.5E-37 | 407 | 521 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 9.42E-24 | 542 | 720 | No hit | No description |
| SuperFamily | SSF55961 | 9.42E-24 | 749 | 782 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 796 aa Download sequence Send to blast |
MSFGGFLDNS TGSGGGARIV ADISYNNTSS STHSNNMPSS ALAQPRLVTQ SLTKSMFNSP 60 GLSLALQTNA DGQGDVTRMA ENFETNVGRR SREEEHESRS GSDNMDGGSG DDQDAADNTN 120 PRKKKRYHRH TPQQIQELEA LFKECPHPDE KQRLELSRRL CLETRQVKFW FQNRRTQMKT 180 QLERHENSLL RQENDKLRAE NMSIRDAMRN PICSNCGGPA IIGEISLEEQ HLRIENARLK 240 DELDRVCALA GKFLGRPISS LATSMGPPLP SSTLELGVGS NGFGGLSSVA TSMPVGPDFG 300 GGIGSAMSVV PHSRPSVTGL DRSMERSMFL ELALAAMDEL VKLAQTDEPL WLRSLEGGRE 360 VLNHEEYMRS FTPCIGLKPN GFVTEASRET GMVIINSLAL VETLMESMHA ELQVLSPLVP 420 VREVNFLRFC KQHAEGVWAV VDVSVDTIRD TSGAPTFMNC RRLPSGCVVQ DMPNGYSKVT 480 WVEHAEYDES QVHQLYRPML SSGMGFGAQR WVATLQRQCE CLAILMSSSV PTRDHTAITA 540 SGRRSMLKLA QRMTDNFCAG VCASTVHKWN KLNARNVDED VRVMTRESLD DPGEPPGIVL 600 SAATSVWLPV SPQRLFDFLR DERLRSEWDI LSNGGPMQEM AHIAKGQDPG NCVSLLRARA 660 MNANQSSMLI LQETCIDSAG GLVVYAPVDI PAMHVVMNGG DSAYVALLPS GFAIVPDGPG 720 SRGPMTVKGG GHGSSNGGGG EDATHRVSGS LLTMTFQILV NSLPSAKLTV ESVETVNNLI 780 SCTVQKIKAA LHCES* |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, stems, leaves and floral buds. {ECO:0000269|PubMed:10402424}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
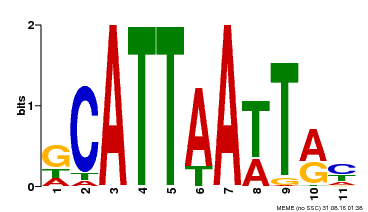
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Prupe.2G156900.2.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007220256.1 | 0.0 | homeobox-leucine zipper protein ANTHOCYANINLESS 2 isoform X1 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A251QJP8 | 0.0 | A0A251QJP8_PRUPE; Uncharacterized protein | ||||
| STRING | EMJ21455 | 0.0 | (Prunus persica) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Prupe.2G156900.2.p |
| Entrez Gene | 18787087 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



