 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pp3c15_4480V3.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Bryophyta; Bryophytina; Bryopsida; Funariidae; Funariales; Funariaceae; Physcomitrella
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 525aa MW: 57094.9 Da PI: 6.7054 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 99.6 | 2.1e-31 | 293 | 348 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k r++W+peLH+rFv+a++qLGGs+ AtPk+i+elmkv+gLt+++vkSHLQkYRl+
Pp3c15_4480V3.1.p 293 KSRRCWSPELHRRFVSALHQLGGSQIATPKQIRELMKVDGLTNDEVKSHLQKYRLH 348
68****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 13.06 | 289 | 350 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.19E-17 | 291 | 351 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 1.0E-28 | 291 | 351 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.0E-25 | 293 | 348 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.4E-7 | 295 | 346 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009266 | Biological Process | response to temperature stimulus | ||||
| GO:0009740 | Biological Process | gibberellic acid mediated signaling pathway | ||||
| GO:0010452 | Biological Process | histone H3-K36 methylation | ||||
| GO:0048579 | Biological Process | negative regulation of long-day photoperiodism, flowering | ||||
| GO:1903507 | Biological Process | negative regulation of nucleic acid-templated transcription | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 525 aa Download sequence Send to blast |
MGSPVDLTLG CSTQAPSSNG YYNHQNEPTH SENYLESLRE VEERVRSLEE ERRKVEGFKR 60 ELPLCIQLLE DAIKTWKVKL VNGKHLPLQL RSGSMDSGRN QHVMPSSPGS RPTLEDFMPF 120 KRRWEQRDQP SESSDVEEEV RGSKRPAWLT EAPLWRQSSG CSEREGKEVQ EHSEFGISER 180 EQSSITSSQM LLSAKQRPAG AFLPFIRDQP GASSFLSRPT PRTVASAADL SLSLGERTTT 240 PSSRNLRPSD SEVGSIDAGL QTPEVQIPKD IGVMSNGTSG AGSSSGGGNP TRKSRRCWSP 300 ELHRRFVSAL HQLGGSQIAT PKQIRELMKV DGLTNDEVKS HLQKYRLHTR RPSSSPQSGG 360 MGQSPQLVVL GGIWVPPQYA ASGSQPSPGF YNPSAAHSPQ PNQFCPTSLP QGYSCISNST 420 QGAQLEMHRE PIFEPPRQAG TSQSQSSPQG PLQVTSQLSG ATQGPSSAAY HDESAGEEDA 480 KSESASWKCD NPKGEELKKS NSNQGSGDGQ SRAHVSEEDV NQAD* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 6e-14 | 293 | 346 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 6e-14 | 293 | 346 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 6e-14 | 293 | 346 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 6e-14 | 293 | 346 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Ppa.2540 | 0.0 | protonema | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00261 | DAP | Transfer from AT2G03500 | Download |
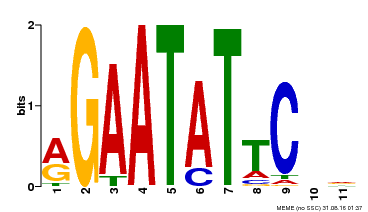
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024395921.1 | 0.0 | myb family transcription factor EFM-like | ||||
| Refseq | XP_024395922.1 | 0.0 | myb family transcription factor EFM-like | ||||
| Refseq | XP_024395923.1 | 0.0 | myb family transcription factor EFM-like | ||||
| TrEMBL | A0A2K1JBV9 | 0.0 | A0A2K1JBV9_PHYPA; Uncharacterized protein | ||||
| STRING | PP1S83_100V6.1 | 0.0 | (Physcomitrella patens) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP4303 | 16 | 24 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G03500.1 | 1e-53 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pp3c15_4480V3.1.p |



