 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_008243945.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Amygdaleae; Prunus
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 701aa MW: 77164.3 Da PI: 6.7818 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 74.3 | 1.4e-23 | 121 | 222 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSS CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrs 88
f k+lt+sd+++ g +++p+ +ae++ ++ + t+ +d +g++W++++iyr++++r++lt+GW+ Fv+ ++L +gD++vF + +
XP_008243945.1 121 FAKTLTQSDANNGGGFSVPRYCAETIfprldysADPPVQ--TILAKDVHGETWKFRHIYRGTPRRHLLTTGWSTFVNHKKLVAGDSIVFL--RAE 211
89****************************996555555..89***********************************************..448 PP
EE..EEEEE-S CS
B3 89 efelvvkvfrk 99
++ l+v+++r+
XP_008243945.1 212 NGDLCVGIRRA 222
8999****997 PP
| |||||||
| 2 | Auxin_resp | 114.2 | 1.2e-37 | 290 | 373 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrl.sGtvvgvsdldpvrWpnSkWrsLk 83
aa+ as++++FevvY+Prast+eF+vk++ v++al++++++GmRfkmafetedss++++ +Gt+++v+ ++p+rWp+S+Wr+L+
XP_008243945.1 290 AATLASNGQPFEVVYYPRASTPEFCVKASLVKAALQIRWCPGMRFKMAFETEDSSRISWfMGTISSVQLAEPLRWPESPWRLLQ 373
6899******************************************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 3.0E-37 | 115 | 225 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 6.41E-29 | 117 | 224 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 2.49E-21 | 120 | 221 | No hit | No description |
| Pfam | PF02362 | 8.8E-22 | 121 | 222 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 13.112 | 121 | 223 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 1.7E-21 | 121 | 223 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 1.2E-32 | 290 | 373 | IPR010525 | Auxin response factor |
| PROSITE profile | PS51745 | 22.3 | 615 | 695 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007389 | Biological Process | pattern specification process | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0048829 | Biological Process | root cap development | ||||
| GO:0051301 | Biological Process | cell division | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 701 aa Download sequence Send to blast |
MITFMDSKEK LEEGEKCLDP QLWHACAGGM VQMPPVNAKV FYFPQGHAEH ACGPVDFRNC 60 PRVPPYIFCR VSAIKFMADP ETDEVYAKIR LVPSSASEAG YEDDGIGGLN GSETPDKPAS 120 FAKTLTQSDA NNGGGFSVPR YCAETIFPRL DYSADPPVQT ILAKDVHGET WKFRHIYRGT 180 PRRHLLTTGW STFVNHKKLV AGDSIVFLRA ENGDLCVGIR RAKRGIGGGP ESSSGWNPTG 240 GNCTMPYGGY SAFLRDDENK LMRNGNGNGS NSNGSLMGKG KVRPESVFEA ATLASNGQPF 300 EVVYYPRAST PEFCVKASLV KAALQIRWCP GMRFKMAFET EDSSRISWFM GTISSVQLAE 360 PLRWPESPWR LLQVTWDEPD LLQNVKRVSP WLVELVSNMP AIHLTPFSPP RKKMRLPQHP 420 DFPFEGQLPM PTFSGNLLGP SSPFGCLPDK TPAGMQGARH GHYGLSLSDM HLNKLQTGLF 480 PASFTPLDHA ATATKFSNNT MTQKPTMSEN VSCLLTMAHS TQTSKKPDDV KPPQLVLFGK 540 PILTEQQISL SCSGDTVSPV LTGNSSSDGN AEKTTNLSDN SGSALHQQSL QERSYCEGFQ 600 WYKVTRQETE SSLETGHCKV FMESEDVGRT LDLSVFGSYD ELYRKLADMF GIENSETLNH 660 VLYRDATGAV KHIGDEPLSD FMKTARRLTI LMDSGSNNVG M |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldu_A | 4e-81 | 18 | 394 | 51 | 388 | Auxin response factor 5 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00461 | DAP | Transfer from AT4G30080 | Download |
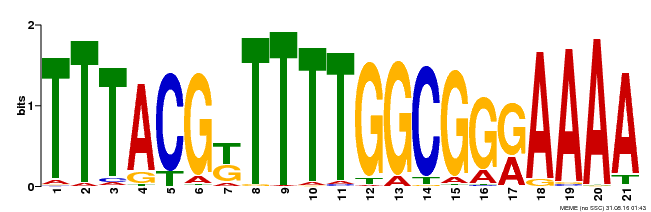
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_008243945.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | FJ177422 | 0.0 | FJ177422.1 Malus x domestica putative auxin response factor ARF16 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008243945.1 | 0.0 | PREDICTED: auxin response factor 18-like isoform X2 | ||||
| Swissprot | Q653H7 | 0.0 | ARFR_ORYSJ; Auxin response factor 18 | ||||
| TrEMBL | A0A314ZP65 | 0.0 | A0A314ZP65_PRUYE; Auxin response factor | ||||
| STRING | XP_008243942.1 | 0.0 | (Prunus mume) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G30080.1 | 0.0 | auxin response factor 16 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



