 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_008233790.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Amygdaleae; Prunus
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 360aa MW: 40640.1 Da PI: 7.2819 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 101.4 | 5.9e-32 | 77 | 130 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
rlrWtp+LH +Fv+ave+LGG+e+AtPk +l+lm+vkgL+++hvkSHLQ+YR+
XP_008233790.1 77 SRLRWTPDLHLCFVHAVERLGGQERATPKLVLQLMNVKGLSIAHVKSHLQMYRS 130
69***************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 12.847 | 73 | 133 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 5.7E-30 | 73 | 131 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 6.81E-15 | 74 | 131 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 4.5E-23 | 77 | 132 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 3.8E-9 | 78 | 129 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 360 aa Download sequence Send to blast |
MKSKSSTEEV SETSPSISKK EFEDEDDEGD EGDEEEMDKD GLQLRNNGVS SSNSTIEENE 60 KKGVSGSVRQ YVRSKTSRLR WTPDLHLCFV HAVERLGGQE RATPKLVLQL MNVKGLSIAH 120 VKSHLQMYRS KKMEDPDQVM SEQGFFMEGG DNHIYNLTQL SMLQSFNQWP SSGLRYGTAT 180 DASWRGRQHQ IYRPYNTHRT ALLDHNTRNI GLYGSVADRI FGNNNKNNPT SANNHLHTNY 240 PSPSYVQTTW RRHQIRDESF QSMINVPCRD HSWREIRGGQ NMLKRKAPDS DHTTCDLDLN 300 LSLKVPPKDD HEFGKGLEGC SLSLSLSSSS SSKLSMKKLE GGGHGRGEHG RMPSTLDLTL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 6e-18 | 77 | 132 | 2 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 6e-18 | 77 | 132 | 2 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 6e-18 | 77 | 132 | 2 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 6e-18 | 77 | 132 | 2 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00301 | DAP | Transfer from AT2G40260 | Download |
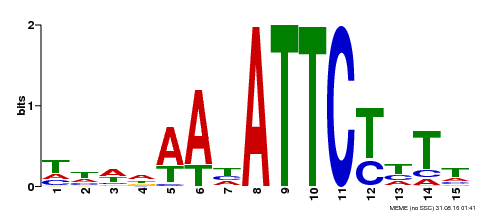
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_008233790.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008233790.1 | 0.0 | PREDICTED: uncharacterized protein LOC103332813 | ||||
| TrEMBL | A0A314ZEH4 | 0.0 | A0A314ZEH4_PRUYE; Uncharacterized protein | ||||
| STRING | XP_008233790.1 | 0.0 | (Prunus mume) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF2378 | 34 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G40260.1 | 5e-49 | G2-like family protein | ||||



