 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_008232742.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Amygdaleae; Prunus
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 828aa MW: 90119.1 Da PI: 6.3958 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 65.4 | 8e-21 | 123 | 178 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r eL+++l L++rqVk+WFqNrR+++k
XP_008232742.1 123 KKRYHRHTPQQIQELEALFKECPHPDEKQRLELSRRLCLETRQVKFWFQNRRTQMK 178
688999***********************************************999 PP
| |||||||
| 2 | START | 206.7 | 9.1e-65 | 330 | 554 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE....EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEEE CS
START 1 elaeeaaqelvkkalaeepgWvkss....esengdevlqkfeeskv.....dsgealrasgvvdmvlallveellddkeqWdetla....kaetl 82
ela++a++elvk+a+ +ep+W +s e +n +e++++f++ + + +ea+r++g+v+ ++ lve+l++++ +W e+++ + +t+
XP_008232742.1 330 ELALAAMDELVKLAQTDEPLWLRSLeggrEVLNHEEYMRSFTPCIGlkpngFVTEASRETGMVIINSLALVETLMESN-RWLEMFPclvaRTSTT 423
5899************************99***********9999899999***************************.**************** PP
EEECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--.-TTSEE-EESSEEEEEEEECTCEEEEEEEE-E CS
START 83 evissg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppesssvvRaellpSgiliepksnghskvtwvehv 170
+vissg galqlm aelq+lsplvp R++ f+R+++q+ +g+w++vdvSvd ++ + + ++ +++lpSg+++++++ng+skvtwveh+
XP_008232742.1 424 DVISSGmggtrnGALQLMHAELQVLSPLVPvREVNFLRFCKQHAEGVWAVVDVSVDTIRDTSGAPTFMNCRRLPSGCVVQDMPNGYSKVTWVEHA 518
************************************************************999******************************** PP
E--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 171 dlkgrlphwllrslvksglaegaktwvatlqrqcek 206
+++++++h+l+r++++sg+ +ga++wvatlqrqce+
XP_008232742.1 519 EYDESQVHQLYRPMLSSGMGFGAQRWVATLQRQCEC 554
**********************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 4.23E-21 | 103 | 180 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.7E-22 | 107 | 180 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.524 | 120 | 180 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 5.7E-17 | 121 | 184 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 2.25E-18 | 123 | 180 | No hit | No description |
| Pfam | PF00046 | 2.2E-18 | 123 | 178 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 155 | 178 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 43.985 | 321 | 557 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 8.79E-33 | 322 | 554 | No hit | No description |
| CDD | cd08875 | 2.60E-125 | 325 | 553 | No hit | No description |
| SMART | SM00234 | 8.6E-47 | 330 | 554 | IPR002913 | START domain |
| Pfam | PF01852 | 8.8E-57 | 330 | 554 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.73E-23 | 575 | 753 | No hit | No description |
| SuperFamily | SSF55961 | 1.73E-23 | 782 | 815 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 828 aa Download sequence Send to blast |
MSFGGFLDNS TGSGGGARIV ADISYNNTSS STHSNNMPSS ALAQPRLVTQ SLTKSMFNSP 60 GLSLALTNAD GQGDVTRMAE NFETHVGRRS REEEHESRSG SDNMDGGSGD DQDAADNTNP 120 RKKKRYHRHT PQQIQELEAL FKECPHPDEK QRLELSRRLC LETRQVKFWF QNRRTQMKTQ 180 LERHENSLLR QENDKLRAEN MSIRDAMRNP ICSNCGGPAI IGEISLEEQH LRIENARLKD 240 ELDRVCALAG KFLGRPISSL ATSMGPPLPS STLELGVGSN GFGGLSSVAT SMPVGPDFGG 300 GIGSAMSVVP HSRPSATGLD RSMERSMFLE LALAAMDELV KLAQTDEPLW LRSLEGGREV 360 LNHEEYMRSF TPCIGLKPNG FVTEASRETG MVIINSLALV ETLMESNRWL EMFPCLVART 420 STTDVISSGM GGTRNGALQL MHAELQVLSP LVPVREVNFL RFCKQHAEGV WAVVDVSVDT 480 IRDTSGAPTF MNCRRLPSGC VVQDMPNGYS KVTWVEHAEY DESQVHQLYR PMLSSGMGFG 540 AQRWVATLQR QCECLAILMS SSVPTRDHTA ITSSGRRSML KLAQRMTDNF CAGVCASTVH 600 KWNKLNARNV DEDVRVMTRE SLDDPGEPPG IVLSAATSVW LPVSPQRLFD FLRDERLRSE 660 WDILSNGGPM QEMAHIAKGQ DPGNCVSLLR ARAMNANQSS MLILQETCID AAGGLVVYAP 720 VDIPAMHVVM NGGDSAYVAL LPSGFAIVPD GPGSRGPMTV KGGGHGSNNG GGGEDATHRV 780 SGSLLTMTFQ ILVNSLPSAK LTVESVETVN NLISCTVQKI KAALHCES |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
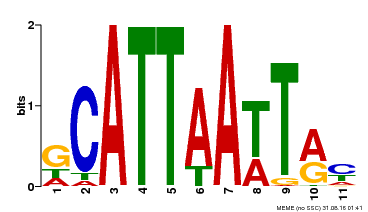
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_008232742.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008232742.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ANTHOCYANINLESS 2 isoform X2 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | M5XQZ7 | 0.0 | M5XQZ7_PRUPE; Uncharacterized protein | ||||
| STRING | XP_008232741.1 | 0.0 | (Prunus mume) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



