 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_008223487.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Amygdaleae; Prunus
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 997aa MW: 110845 Da PI: 6.5069 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 130.9 | 4.5e-41 | 142 | 219 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqve+C+adls+ak+yhrrhkvC++hska+++lv + +qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+++
XP_008223487.1 142 VCQVEDCKADLSNAKDYHRRHKVCDMHSKASTALVGNAMQRFCQQCSRFHVLQEFDEGKRSCRRRLAGHNRRRRKTHP 219
6**************************************************************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 7.2E-33 | 137 | 204 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.052 | 140 | 217 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.18E-37 | 141 | 222 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 2.7E-29 | 143 | 216 | IPR004333 | Transcription factor, SBP-box |
| SMART | SM00248 | 150 | 765 | 794 | IPR002110 | Ankyrin repeat |
| Gene3D | G3DSA:1.25.40.20 | 1.5E-7 | 767 | 880 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.27E-9 | 767 | 879 | No hit | No description |
| SuperFamily | SSF48403 | 1.27E-8 | 767 | 880 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 3.5 | 820 | 850 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 4200 | 860 | 890 | IPR002110 | Ankyrin repeat |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 997 aa Download sequence Send to blast |
MEAEFGGKAH SYYGMKAVGK KSFEWDLNDW KWDGDLFTAS PLNSVPSACR SKQLFPVRPE 60 TPSNAGLSNS SSSGSDNISP GNEKGKRELE KRRRAVFVEN EVHDEAGSLN LNLGGQAYPI 120 MEGEVQTGKK TKIVGTTSNC AVCQVEDCKA DLSNAKDYHR RHKVCDMHSK ASTALVGNAM 180 QRFCQQCSRF HVLQEFDEGK RSCRRRLAGH NRRRRKTHPD TTANGGSLND ERGSSYLLIS 240 LLRILSNMHS SSSDQTKDQD LLSHLLRSLA NLAGTADGRN ISTLLHGSQG LFNSGTSVQT 300 TKVPDMDDGV NLEDLRPVGQ CSVVPASDML ERRISSVDDP GSLQVLSGLQ ATEPLPSRDS 360 SESKSVTPEA TSRRFQLNGI DLNNTYDDSQ EYLENLGNSH VPASPGTASW LQRDSHKSSP 420 PQTSGNSDLT STQSPSSSSG EAQSRTDRIV FKLFGKDPND LPFILRSQIL DWLSHSPTDI 480 ESYIRPGCII LTIYLRLENS TWEELCCHLG SSLKTLLDAA DDPFWRTGWV YTRVQHFVTF 540 TYNGQVVLDT PLPLKSDKSC RISYIKPIAV SVSERAQFVV KGFNLSHSAT RLLCALEGKY 600 LVQETCYDMM DGVHTTVEHD ELQCLKFSCS IPDVTGRGFI EVEDHGLSSS FFPFIVAEQE 660 VCSEICMLED EIEVPESADA EKLEAKNQAL DFIHELGWLL HRSRAKFRLG HSDLNLDLFP 720 FSRFRLLMEF SIEHDWCVVV KKLLGILFEG TVDAGEHTSV EFALLDMSLL HRAVQRNCRS 780 MVEFLLKFIP NQGLTGSEQK QQVDRDGNSF LFKPDAVGPM GLTPLHVAAS ADGYEHVLDA 840 LTDDPGKLGI EAWKNARDST GLTPYDYACL QSRYSYVHLV QRKISKTLES GQVVLDIPGV 900 ILDRNGKQKQ SEAYKPSRVA SLETEKIEMK VILRHCKLCA QKPAYGNTRS LVYRPAMLSM 960 VAVAAVCVCV ALLFKSTPEV LFVFQPFRWE LLKYGSS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 6e-30 | 142 | 216 | 10 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' of AP1 promoter. Binds specifically to the 5'-GTAC-3' core sequence. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:16095614}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00603 | PBM | Transfer from AT2G47070 | Download |
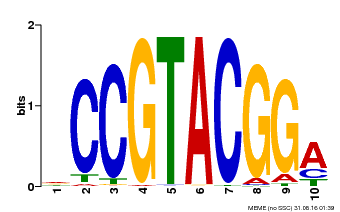
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_008223487.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KJ622309 | 8e-65 | KJ622309.1 Gossypium hirsutum SQUAMOSA promoter binding-like transcription factor (SPL1) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008223487.1 | 0.0 | PREDICTED: squamosa promoter-binding-like protein 1 | ||||
| Swissprot | Q9SMX9 | 0.0 | SPL1_ARATH; Squamosa promoter-binding-like protein 1 | ||||
| TrEMBL | M5XKY4 | 0.0 | M5XKY4_PRUPE; Uncharacterized protein | ||||
| STRING | XP_008223487.1 | 0.0 | (Prunus mume) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1539 | 34 | 95 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



