 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PSME_00054096-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Acrogymnospermae; Pinidae; Pinales; Pinaceae; Pseudotsuga
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 395aa MW: 43958.6 Da PI: 5.0151 | ||||||||
| Description | GATA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GATA | 55 | 1.1e-17 | 270 | 304 | 1 | 35 |
GATA 1 CsnCgttkTplWRrgpdgnktLCnaCGlyyrkkgl 35
C +C t kTp+WR gp g+ktLCnaCG++y++ +l
PSME_00054096-RA 270 CMHCDTMKTPQWRTGPMGPKTLCNACGVRYKSGRL 304
99*****************************9885 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50114 | 11.517 | 264 | 300 | IPR000679 | Zinc finger, GATA-type |
| SMART | SM00401 | 2.3E-15 | 264 | 314 | IPR000679 | Zinc finger, GATA-type |
| SuperFamily | SSF57716 | 7.13E-15 | 268 | 327 | No hit | No description |
| Gene3D | G3DSA:3.30.50.10 | 1.2E-14 | 268 | 302 | IPR013088 | Zinc finger, NHR/GATA-type |
| CDD | cd00202 | 1.10E-12 | 269 | 316 | No hit | No description |
| PROSITE pattern | PS00344 | 0 | 270 | 295 | IPR000679 | Zinc finger, GATA-type |
| Pfam | PF00320 | 4.3E-15 | 270 | 304 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009416 | Biological Process | response to light stimulus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 395 aa Download sequence Send to blast |
MEAADCLYGT NYGTPFLDGK KSSGKFTELK NGVQLQDFFH IDDLLDFSNE EIAGPIGDDG 60 NNWPSCTTTA VTAAADSSTI TLTETLSCNS SLDGKSFRAY NVLETDENNN GHLCVPCDDL 120 AELEWLSNFV EDSFSTEQAP KPIFPSNFGD VRTVKEEEKP HIAIPIRSIS PMSGLDTSSS 180 SWKTPSFCPE TPVPGRARSK RSRAPACNWS SRIVSPASSA SALENGPTSS VLSSDSEFFA 240 ESYFPAKKPA KMFHGNKKRE QEQLTQIRKC MHCDTMKTPQ WRTGPMGPKT LCNACGVRYK 300 SGRLVPEYRP AASPTFVVSK HSNSHRKVIE MRRQKELQEA LQQASKLYSN DKQGDEEYGD 360 EYEEEEDGSD GQYPHFGSDD DNYLVHTKQD YRHFI |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00048 | PBM | Transfer from AT4G32890 | Download |
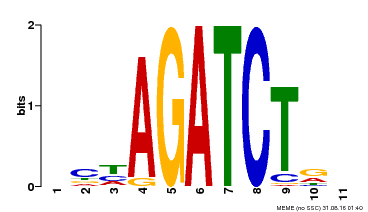
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT107012 | 0.0 | BT107012.1 Picea glauca clone GQ03101_M09 mRNA sequence. | |||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G32890.1 | 1e-59 | GATA transcription factor 9 | ||||



