 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PSME_00050134-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Acrogymnospermae; Pinidae; Pinales; Pinaceae; Pseudotsuga
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 425aa MW: 47508.9 Da PI: 4.6796 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 56.6 | 6.4e-18 | 100 | 149 | 2 | 55 |
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
++ GVr+++ +grWvAeI+d + ++ r +lg+f taeeAa+a+++a+ l+g
PSME_00050134-RA 100 RFVGVRQRP-SGRWVAEIKDTT---QKIRMWLGTFETAEEAARAYDEAACLLRG 149
789******.7*********32...35**********************99987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 2.77E-31 | 99 | 156 | No hit | No description |
| Gene3D | G3DSA:3.30.730.10 | 1.8E-27 | 100 | 156 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 9.15E-19 | 100 | 157 | IPR016177 | DNA-binding domain |
| SMART | SM00380 | 3.1E-36 | 100 | 163 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.585 | 100 | 157 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 1.4E-12 | 100 | 149 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.9E-7 | 101 | 112 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.9E-7 | 123 | 139 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000302 | Biological Process | response to reactive oxygen species | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0035865 | Biological Process | cellular response to potassium ion | ||||
| GO:0048528 | Biological Process | post-embryonic root development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 425 aa Download sequence Send to blast |
MDSLPSFTMS ASNGSSLTHP YSLRSASLQS SNCFNKPEAC AAPTAQFLAM AVQPSANTNH 60 VNANSNNLHA NFNPPITTGF DQSSGAMQTM MKNSNSRRKR FVGVRQRPSG RWVAEIKDTT 120 QKIRMWLGTF ETAEEAARAY DEAACLLRGS NTRTNFVPTV NSCKNSALAS RINRLLSLRK 180 DSTASANSKS IIKEDCKDRH DDADDNQNDS FKSSTNYDVS EDSNEDSLPI EQSPSCTSSF 240 SNYVCSSSSS SSSAMDDDMD QASLRQQNGE KQPHFSHDIH TQHGQIYSSV TEEEEMKNQS 300 RSILTFNLSN FDKNHFIDPI SIEEEMINQS RSISTPDLFN TNKEEERNHF SMEDEMSGYS 360 CFSSSAEYFW PLDFGLDIQF PVDEFPVIDD LPALSMSDYH DHARASCHVQ PEFCPQSDYL 420 SDLIR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2gcc_A | 1e-15 | 96 | 162 | 1 | 68 | ATERF1 |
| 3gcc_A | 1e-15 | 96 | 162 | 1 | 68 | ATERF1 |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00521 | DAP | Transfer from AT5G19790 | Download |
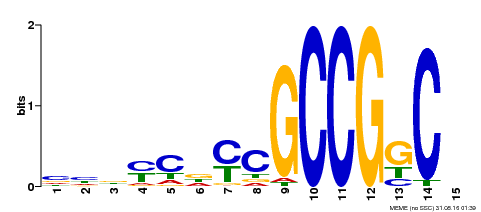
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G19790.1 | 2e-24 | related to AP2 11 | ||||



