 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PSME_00021294-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Acrogymnospermae; Pinidae; Pinales; Pinaceae; Pseudotsuga
|
||||||||
| Family | RAV | ||||||||
| Protein Properties | Length: 620aa MW: 69367.5 Da PI: 7.0171 | ||||||||
| Description | RAV family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 43.1 | 1.1e-13 | 175 | 223 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s+ykGV + +grW A+I++ ++r++lg+f +e+Aaka+++a++k++g
PSME_00021294-RA 175 SHYKGVVPQP-NGRWGAQIYEK-----HQRVWLGTFNKEEDAAKAYDRAALKFRG 223
79****9888.8*********3.....5*************************98 PP
| |||||||
| 2 | B3 | 100.1 | 1.3e-31 | 328 | 431 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SS CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgr 87
f+k+ tpsdv+k++rlv+pk++ae++ + + ++ l +ed +g+ W+++++y+++s++yvltkGW++Fvk+++L++gD+v F++ +
PSME_00021294-RA 328 FDKAVTPSDVGKLNRLVIPKQHAEKYfpldvnS--SDKGLLLNFEDNTGKAWRFRYSYWNSSQSYVLTKGWSRFVKDKKLEAGDIVSFQRGLQ 418
89***************************9643..447789************************************************8844 PP
.SEE..EEEEE-S CS
B3 88 .sefelvvkvfrk 99
++++l+++++r+
PSME_00021294-RA 419 aQSEQLYISWRRR 431
499999****998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 8.91E-26 | 175 | 230 | No hit | No description |
| SuperFamily | SSF54171 | 1.5E-17 | 175 | 231 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 1.3E-8 | 175 | 223 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 2.5E-29 | 176 | 237 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 20.323 | 176 | 231 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 3.1E-21 | 176 | 231 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:2.40.330.10 | 7.8E-40 | 319 | 432 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 1.06E-28 | 324 | 416 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 1.75E-25 | 326 | 415 | No hit | No description |
| SMART | SM01019 | 2.7E-22 | 328 | 432 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 1.2E-28 | 328 | 431 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 15.016 | 328 | 432 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009910 | Biological Process | negative regulation of flower development | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0048527 | Biological Process | lateral root development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 620 aa Download sequence Send to blast |
MMDLQGRGKR SAATTSLWQS EIDSPHVELS LGASCQASDV ETIICSQKNA SASTPTIHVL 60 GRQDQYARVE AQEHTTNEAM KDQEDNQDDE MGSAMTEEFN YSVMAVSENE KQPKLEAALF 120 KDWCSARSME KNLQGLDRMG SGISVLSVSG GESKGFDDLR RYSMPMDQAG KLPSSHYKGV 180 VPQPNGRWGA QIYEKHQRVW LGTFNKEEDA AKAYDRAALK FRGRDAMTNF SPVEDSDPEA 240 RFMNEHNKRQ IVDMLRKHTY DEELDHHSKR FKMIAARSVA MNNAEVSGLG AHDVCQENQV 300 SVDENMNAAS SSTAPVADRA GLPREHLFDK AVTPSDVGKL NRLVIPKQHA EKYFPLDVNS 360 SDKGLLLNFE DNTGKAWRFR YSYWNSSQSY VLTKGWSRFV KDKKLEAGDI VSFQRGLQAQ 420 SEQLYISWRR RPPCQPRLGV GTQSLQSKGG SVYPVKNLSF LHPFNSGTPV VYDAQWMPIF 480 WPSSTNPDAF TQLNSHFVSP IVNKVDALWS QSPVTTLNGA LPFYFPASPT NQMLQSRFYN 540 NNSDGCQPMV DLVNQHNSIN PCNGNTNIKP LHPFVNAARP YSADELETPE HKSASSLDPA 600 RKKGVRLFGV TLSELPHRLS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5os9_A | 1e-56 | 324 | 432 | 4 | 114 | B3 domain-containing transcription factor NGA1 |
| 5os9_B | 1e-56 | 324 | 432 | 4 | 114 | B3 domain-containing transcription factor NGA1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00139 | DAP | Transfer from AT1G13260 | Download |
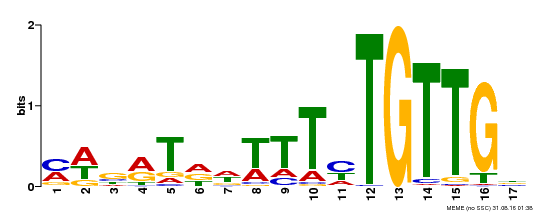
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| TrEMBL | A0A0D6QRH4 | 1e-117 | A0A0D6QRH4_ARACU; Uncharacterized protein | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G13260.1 | 8e-98 | related to ABI3/VP1 1 | ||||



