 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PSME_00007067-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Acrogymnospermae; Pinidae; Pinales; Pinaceae; Pseudotsuga
|
||||||||
| Family | CPP | ||||||||
| Protein Properties | Length: 772aa MW: 82927 Da PI: 8.4859 | ||||||||
| Description | CPP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCR | 49.6 | 7.7e-16 | 303 | 341 | 3 | 41 |
TCR 3 kkgCnCkkskClkkYCeCfaagkkCseeCkCedCkNkee 41
+k+CnCkkskClk+YCeCfaag++C e C+C++C+Nk e
PSME_00007067-RA 303 CKRCNCKKSKCLKLYCECFAAGVYCVEPCTCQECFNKPE 341
79**********************************976 PP
| |||||||
| 2 | TCR | 50.3 | 4.6e-16 | 387 | 425 | 1 | 39 |
TCR 1 kekkgCnCkkskClkkYCeCfaagkkCseeCkCedCkNk 39
++k+gCnCkks ClkkYCeC++ag+ Cs+ C+Ce+CkN
PSME_00007067-RA 387 RHKRGCNCKKSMCLKKYCECYQAGVGCSDGCRCESCKNV 425
589***********************************6 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01114 | 8.2E-16 | 301 | 342 | IPR033467 | Tesmin/TSO1-like CXC domain |
| PROSITE profile | PS51634 | 35.511 | 302 | 427 | IPR005172 | CRC domain |
| Pfam | PF03638 | 1.2E-11 | 304 | 339 | IPR005172 | CRC domain |
| SMART | SM01114 | 1.5E-16 | 387 | 428 | IPR033467 | Tesmin/TSO1-like CXC domain |
| Pfam | PF03638 | 8.1E-12 | 389 | 424 | IPR005172 | CRC domain |
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 772 aa Download sequence Send to blast |
MQPGNQSQRG IRRRCLDFEA SEAQRKTMSN SSWKSRSQMP KTGALVNSTH DSETNTSPSV 60 SCEKGVTSDC KQIVPLKRET GVDSGRNSQS SSNVRFSTFL SKTNELTSDK SDSNVRNNGN 120 TPISVGIPSG IGLHLNSLAT AMPLNCGVNM LTSAKGSAST QGMSPSSGVK EASSELVGSS 180 LPVVNSGGNI PVGSNSNIAS GMVVSMVEKT RIDHQELQSS GMSQIGVTRS TTCFRSVTVG 240 IKPLQSRLPF KSEERNLSPL GKKRPALLDI SQQSPFSTGE EFSQSSPKKK RRKSTTIGDK 300 DGCKRCNCKK SKCLKLYCEC FAAGVYCVEP CTCQECFNKP EYEDMVLGTR QQIESRNPLA 360 FAPKIVRGAD SPPTNGDECG ETPSSARHKR GCNCKKSMCL KKYCECYQAG VGCSDGCRCE 420 SCKNVYGKKE GGSDDIEDNE VPIEGWDKDS LEEKAEVHDV GNDILLSEQQ HANDLSPLTP 480 SFQYSGQGKS SAKLNSCGKK HFASEDLESP TVSQSCAKPP RSPGKILRPT KGIQGNITAI 540 HNRQAGSRTS ASPMFTPKMD KSGKFSPQWD CLADICTLTP MLHPPMRPSA SSASNVDGID 600 VSPFSGQQNE ISSMSSRPLA SRHSCHVGTS IGFRQPAARS PICTSENIHW QTPVNAKVTP 660 VTPALSTTAC TAGNKLSDGS DFDMQSHDVS DSLEDDTPDI LKNNGSPTRG LKASSPNQKR 720 VSPPHNYCPK ELMNMRPISS TPGIRSGRKY ILQSVPSFPP LTPLSGEGQA NE |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5fd3_A | 6e-18 | 306 | 425 | 14 | 121 | Protein lin-54 homolog |
| 5fd3_B | 6e-18 | 306 | 425 | 14 | 121 | Protein lin-54 homolog |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 287 | 291 | KKKRR |
| 2 | 287 | 292 | KKKRRK |
| 3 | 288 | 292 | KKRRK |
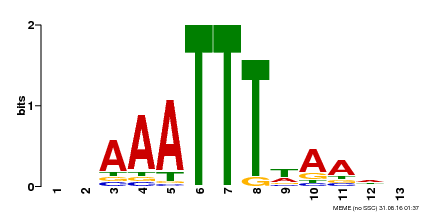
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00624 | PBM | Transfer from PK22848.1 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT123813 | 1e-134 | BT123813.1 Picea sitchensis clone WS04717_J09 unknown mRNA. | |||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G14770.1 | 3e-69 | TESMIN/TSO1-like CXC 2 | ||||



