 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Peinf101Scf00175g04023.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Petunioideae; Petunia; Petunia integrifolia
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 812aa MW: 88959.3 Da PI: 6.3007 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 65.6 | 6.8e-21 | 107 | 162 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r eL+k+l L++rqVk+WFqNrR+++k
Peinf101Scf00175g04023.1 107 KKRYHRHTPQQIQELEALFKECPHPDEKQRLELSKRLCLETRQVKFWFQNRRTQMK 162
678899***********************************************999 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 4.4E-22 | 93 | 164 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 1.03E-20 | 95 | 164 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.54 | 104 | 164 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 3.1E-17 | 105 | 168 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.30E-18 | 106 | 164 | No hit | No description |
| Pfam | PF00046 | 2.0E-18 | 107 | 162 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 139 | 162 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 44.034 | 307 | 544 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 2.2E-33 | 309 | 541 | No hit | No description |
| CDD | cd08875 | 2.98E-123 | 311 | 540 | No hit | No description |
| SMART | SM00234 | 1.9E-50 | 316 | 541 | IPR002913 | START domain |
| Pfam | PF01852 | 2.9E-57 | 316 | 541 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 5.77E-24 | 560 | 798 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 812 aa Download sequence Send to blast |
MFNSPGLSLA LVSHKNLIFT LSSIGHGFSY ISPFADLSGK YVVVQTGMEG GQSEVTRMPE 60 TYLEGNNSNG RRSREEEPES RSGSDNLEEG ASGDDQEDAA DKPPRRKKRY HRHTPQQIQE 120 LEALFKECPH PDEKQRLELS KRLCLETRQV KFWFQNRRTQ MKTQLERHEN SLLRQENDKL 180 RAENMSIREA MRNPICTNCG GPAMIGEISL EEQHLRIENA RLKDELDRVC GLAGKFLGRP 240 ISSLVTSMPP PMPNSSLELG VGNNGFVVGG LSNVPTTLPL APSPDFGVGI ANSLPMVTRQ 300 ASTGIERSLE RSMYLELALA AMDELVKLAQ TDEPLWFRSM EGGGKEILNH EEYMRTFTPC 360 IGMRPNSFVS EASRETGMVI INSMALVETL MDSNKWAEMF PCLIPRTSTT DVISSGMGGT 420 RNGALQLMHA ELQVLSPLVP IREVNFLRFC KQHAEGVWAV VDVSVDNIRE ASGAPTFPNC 480 RRLPSGCVVQ DMPNGYSKVT WVEHAEYDES VIHHLYRSLI SAGMGFGAQR WVATLQRQCE 540 CLAILMSSTV SSRDHTAITP SGRRSMLKLA QRMTNNFCAG VCASTVHKWN KLCAGNVDED 600 VRVMTRKSVD DPGEPPGIVL SAATSVWLPV SPQRLFDFLR DERLRSEWDI LSNGGPMQEM 660 AHIAKGQDHG NCVSLLRASA MNTNQSSMLI LQETCIDAAG ALVVYAPVDI PAMHVVMNGG 720 DSAYVALLPS GFSIVPDGPG SRGPNNANCG SNGPSCNGGP GERISGSLLT VAFQILVNSL 780 PTAKLTVESV ETVNNLISCT VQKIKAALHC ES |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
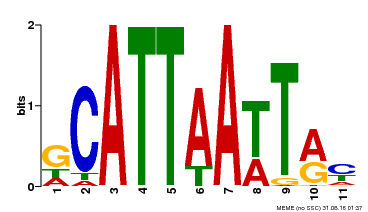
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | GQ222185 | 0.0 | GQ222185.1 Solanum lycopersicum cultivar M82 cutin deficient 2 (CD2) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_019259406.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A1J6ISN2 | 0.0 | A0A1J6ISN2_NICAT; Homeobox-leucine zipper protein anthocyaninless 2 | ||||
| STRING | XP_009782443.1 | 0.0 | (Nicotiana sylvestris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA1626 | 24 | 65 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



