 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Peinf101Scf00145g23014.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Petunioideae; Petunia; Petunia integrifolia
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 489aa MW: 53689.8 Da PI: 6.053 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 52.5 | 1.1e-16 | 37 | 84 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT+ Ed +lvd+++++G g+W+++ ++ g+ R++k+c++rw ++l
Peinf101Scf00145g23014.1 37 KGPWTKAEDAILVDYIARHGEGNWSAVRKHSGLARCGKSCRLRWANHL 84
79******************************************9986 PP
| |||||||
| 2 | Myb_DNA-binding | 51.7 | 2.1e-16 | 90 | 133 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
+g++T+eE+ l++++++++G++ W++ a++++ gRt++++k++w++
Peinf101Scf00145g23014.1 90 KGAFTPEEERLIIELHAKMGNK-WARMAAELP-GRTDNEIKNYWNT 133
799*******************.*********.***********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 17.566 | 32 | 84 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 4.87E-30 | 34 | 131 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 7.1E-14 | 36 | 86 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 9.8E-15 | 37 | 84 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.1E-23 | 38 | 91 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.13E-11 | 39 | 84 | No hit | No description |
| PROSITE profile | PS51294 | 26.27 | 85 | 139 | IPR017930 | Myb domain |
| SMART | SM00717 | 1.3E-16 | 89 | 137 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.3E-15 | 90 | 133 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 9.5E-26 | 92 | 138 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.25E-12 | 92 | 133 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009789 | Biological Process | positive regulation of abscisic acid-activated signaling pathway | ||||
| GO:0043068 | Biological Process | positive regulation of programmed cell death | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0045926 | Biological Process | negative regulation of growth | ||||
| GO:0048235 | Biological Process | pollen sperm cell differentiation | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 489 aa Download sequence Send to blast |
MTSECVDRMT SKVGVDSPAV EEASGGGNAG GGVPLKKGPW TKAEDAILVD YIARHGEGNW 60 SAVRKHSGLA RCGKSCRLRW ANHLRPDLRK GAFTPEEERL IIELHAKMGN KWARMAAELP 120 GRTDNEIKNY WNTRIKRRQR AGMPMYPPDI CYQAFSENKQ NKELGTFSSA DSHPDFSPIN 180 SFEIPAEFKT LELSQQWYPP VLLDIATNSS LDIPATSLLA QGLNFSCNTR SFLSTMHPSK 240 RIRGLESWFS GLNVDFFQAC HQFENDGSLL AESLGYSSPH TDNLISVHHP SFSGVFTGSH 300 APLNGNSSSE PKWAKKLELP SLQTQMVSWG SPSPLPSLES YDTLIQLPPS EHTDSGSLSP 360 GNSGLLDAVL YESQTFRASK NNLHQETCDV VDDSCPDVQA TQWGTHSGPN SPLGHSAASV 420 FSEYTPIFGG SLHEPRSVAS LFGENGCKVK QEDVDLAPTD RNDDVSNHQI FAWTESCFAP 480 TVLVPQQNR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 2e-32 | 32 | 138 | 1 | 107 | B-MYB |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00490 | DAP | Transfer from AT5G06100 | Download |
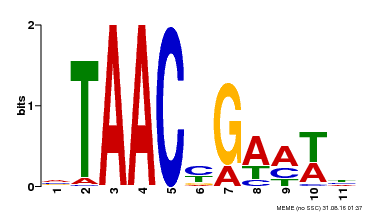
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009771204.1 | 0.0 | PREDICTED: transcription factor GAMYB-like isoform X1 | ||||
| TrEMBL | A0A1U7W971 | 0.0 | A0A1U7W971_NICSY; transcription factor GAMYB-like isoform X1 | ||||
| STRING | XP_009771204.1 | 0.0 | (Nicotiana sylvestris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA2021 | 23 | 58 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G06100.1 | 1e-76 | myb domain protein 33 | ||||



