 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PH01001287G0090 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Bambusoideae; Arundinarodae; Arundinarieae; Arundinariinae; Phyllostachys
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 253aa MW: 28133.7 Da PI: 7.2869 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 50.1 | 6.3e-16 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT+eEd+ lv ++++G +W++ ++ g+ R++k+c++rw +yl
PH01001287G0090 14 KGPWTPEEDKVLVAHIERFGHSNWRALPKLAGLLRCGKSCRLRWINYL 61
79******************************99************97 PP
| |||||||
| 2 | Myb_DNA-binding | 53.7 | 4.7e-17 | 67 | 112 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg++T+eE++ +++++++lG++ W++Ia++++ gRt++++k+ w+++l
PH01001287G0090 67 RGNFTKEEEDTIIQLHELLGNR-WSAIAARLP-GRTDNEIKNVWHTHL 112
89********************.*********.************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 2.9E-24 | 5 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 14.868 | 9 | 61 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 5.14E-31 | 10 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 2.1E-12 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 4.2E-14 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 4.19E-10 | 16 | 61 | No hit | No description |
| PROSITE profile | PS51294 | 25.983 | 62 | 116 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 6.9E-27 | 65 | 116 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 2.0E-15 | 66 | 114 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 4.3E-16 | 67 | 112 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.40E-11 | 69 | 112 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 253 aa Download sequence Send to blast |
MGRAPCCEKM GLKKGPWTPE EDKVLVAHIE RFGHSNWRAL PKLAGLLRCG KSCRLRWINY 60 LRPDIKRGNF TKEEEDTIIQ LHELLGNRWS AIAARLPGRT DNEIKNVWHT HLKKRLEPKA 120 QEHGAASAKK HKPAAPKRES GGRRTDVPTT APVSPERSVL SSVTESSMAS SVAEHGNTGS 180 SAFAKEESFT SGSEDFQIDE SFWSETLSMP LDSHDVSMEP RDAFGASSGA ADDMDFWLRV 240 FIEAGDVQQL PQI |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1h8a_C | 4e-28 | 12 | 116 | 25 | 128 | MYB TRANSFORMING PROTEIN |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator involved in cold stress response (PubMed:14675437, PubMed:20807373, PubMed:22246661). Regulates positively the expression of genes involved in reactive oxygen species (ROS) scavenging such as peroxidase and superoxide dismutase during cold stress. Transactivates a complex gene network that have major effects on stress tolerance and panicle development (PubMed:20807373). {ECO:0000269|PubMed:14675437, ECO:0000269|PubMed:20807373, ECO:0000269|PubMed:22246661}. | |||||
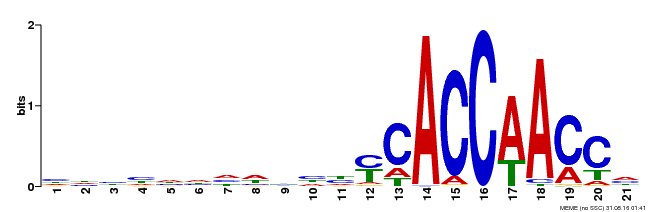
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00375 | DAP | Transfer from AT3G23250 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | PH01001287G0090 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By cold stress. {ECO:0000269|PubMed:14675437, ECO:0000269|PubMed:20807373, ECO:0000269|Ref.2}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | FP093954 | 0.0 | FP093954.1 Phyllostachys edulis cDNA clone: bphyem107b06, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002448188.1 | 1e-123 | transcription factor MYB4 | ||||
| Swissprot | Q7XBH4 | 1e-120 | MYB4_ORYSJ; Transcription factor MYB4 | ||||
| TrEMBL | A0A1D8F0K4 | 0.0 | A0A1D8F0K4_9POAL; MYB protein | ||||
| STRING | Sb06g022660.1 | 1e-122 | (Sorghum bicolor) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP79 | 38 | 563 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G23250.1 | 2e-76 | myb domain protein 15 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



