 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PH01001117G0310 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Bambusoideae; Arundinarodae; Arundinarieae; Arundinariinae; Phyllostachys
|
||||||||
| Family | Dof | ||||||||
| Protein Properties | Length: 318aa MW: 33349.2 Da PI: 8.3669 | ||||||||
| Description | Dof family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-Dof | 117.4 | 5.6e-37 | 75 | 133 | 3 | 61 |
zf-Dof 3 ekalkcprCdstntkfCyynnyslsqPryfCkaCrryWtkGGalrnvPvGggrrknkks 61
e lkcprCdstntkfCy+nnyslsqPr+fC+aCrryWt+GGalrnvPvGgg r++ k+
PH01001117G0310 75 EPGLKCPRCDSTNTKFCYFNNYSLSQPRHFCRACRRYWTRGGALRNVPVGGGYRRHAKR 133
6779*************************************************998765 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| ProDom | PD007478 | 4.0E-35 | 61 | 132 | IPR003851 | Zinc finger, Dof-type |
| PROSITE profile | PS50884 | 28.789 | 78 | 132 | IPR003851 | Zinc finger, Dof-type |
| Pfam | PF02701 | 6.2E-32 | 78 | 132 | IPR003851 | Zinc finger, Dof-type |
| PROSITE pattern | PS01361 | 0 | 80 | 116 | IPR003851 | Zinc finger, Dof-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009640 | Biological Process | photomorphogenesis | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 318 aa Download sequence Send to blast |
MIFPPAFLDS SGWNTNQLQP PTSNNITGPS AVAGGEGNLH QDQDGLATAA GTLPEGGGDG 60 SAMERARLAR IPRPEPGLKC PRCDSTNTKF CYFNNYSLSQ PRHFCRACRR YWTRGGALRN 120 VPVGGGYRRH AKRAKPKATA SATAAASASA VSSTTPSTCT TVAPTLHAML SNNASSLLPP 180 LHRFTDFDTG SLGLSFPGAA KLFADTAAGY AAGGGSSVQG MEQQWRAQAQ MQSFPFLHAM 240 DHQMGPATAM PTAMAMQGMF HLGLESGGGG FNSEDGGEFH VMPTKREEYP RGMYGDVSGY 300 TSYSSSTAGA VTVTIDY* |
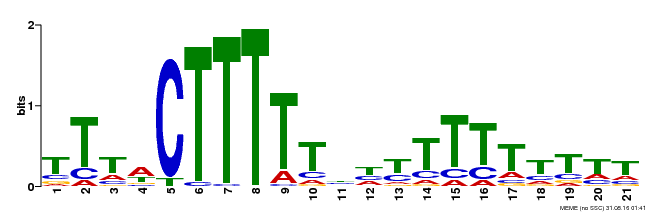
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00407 | DAP | Transfer from AT3G55370 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | PH01001117G0310 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC079356 | 1e-94 | AC079356.2 Oryza sativa Japonica Group cultivar Nipponbare chromosome 5 clone P0016H04, complete sequence. | |||
| GenBank | AK108933 | 1e-94 | AK108933.1 Oryza sativa Japonica Group cDNA clone:002-153-A09, full insert sequence. | |||
| GenBank | AP014961 | 1e-94 | AP014961.1 Oryza sativa Japonica Group DNA, chromosome 5, cultivar: Nipponbare, complete sequence. | |||
| GenBank | GQ183552 | 1e-94 | GQ183552.1 Oryza sativa Japonica Group Dof-type zinc finger protein 25 gene, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015638255.1 | 7e-79 | dof zinc finger protein DOF3.6 isoform X1 | ||||
| TrEMBL | A0A0E0KXP5 | 1e-81 | A0A0E0KXP5_ORYPU; Uncharacterized protein | ||||
| STRING | Pavir.Ca00639.1.p | 2e-83 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP140 | 37 | 367 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G55370.1 | 3e-39 | OBF-binding protein 3 | ||||



