 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PH01000352G0060 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Bambusoideae; Arundinarodae; Arundinarieae; Arundinariinae; Phyllostachys
|
||||||||
| Family | BBR-BPC | ||||||||
| Protein Properties | Length: 380aa MW: 42043.9 Da PI: 10.3174 | ||||||||
| Description | BBR-BPC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GAGA_bind | 400.8 | 1.8e-122 | 51 | 379 | 1 | 301 |
GAGA_bind 1 mdddgsre..rnkg.yye........paaslkenlglqlmssiaerdakirernlalsekkaavaerdmaflqrdkalaernkalverdnklla 83
md+ g+re r+++ +y+ p+++lk++ +++l+++++erd++irer++al+ekkaa+aerdmaf+qrd+a+aern+a+verdn+l+a
PH01000352G0060 51 MDNLGHREngRQRPdQYKavhtqwmmPQRQLKDHHSMNLLALMNERDSAIRERDHALAEKKAAIAERDMAFAQRDAAMAERNAAVVERDNALAA 144
9999*99999****99*99****999889***************************************************************** PP
GAGA_bind 84 lllvensla...salpvgvqvlsgtksidslqqlse.....pqledsave.lreeeklealpieeaaeeakekkkkkkrqrakkpkekkak..k 166
l+l++++ + + + +++lsg+k+i++++qls+ ql+ds+++ +re+++lea+p+++a+ ++ ++k++kk++ +++p+++ + +
PH01000352G0060 145 LELARTNGLnvnNGNEFPQGSLSGAKNIHHHDQLSHvqsspLQLADSPYDhVREMHILEAYPVSTAPGNIGKAKRPKKNNGQASPLKRPSGvlR 238
*999887765667788999**************999999999********9*************************************999887 PP
GAGA_bind 167 kkkksekskkkvkkesader......skaekksidlvlngvslDestlPvPvCsCtGalrqCYkWGnGGWqSaCCtttiSvyPLPvstkrrgaR 254
k+kk ++++k+v +++ e+ +k+e+k++dl+ln+v++Dest+P+P+CsCtG+lrqCYkWGnGGWqS+CCt++iS+yPLPv++++r+aR
PH01000352G0060 239 KTKKATSDWKNVGMSGGGEDsgrvsvMKNEWKDQDLGLNQVAFDESTMPAPACSCTGKLRQCYKWGNGGWQSSCCTMSISMYPLPVMPNKRHAR 332
77778899999999987744688999******************************************************************** PP
GAGA_bind 255 iagrKmSqgafkklLekLaaeGydlsnpvDLkdhWAkHGtnkfvtir 301
++grKmS+gaf+klL++LaaeG+dls+pvDLkdhWAkHGtn+++tir
PH01000352G0060 333 MGGRKMSGGAFTKLLSRLAAEGHDLSTPVDLKDHWAKHGTNRYITIR 379
**********************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF06217 | 1.0E-97 | 51 | 379 | IPR010409 | GAGA-binding transcriptional activator |
| SMART | SM01226 | 1.9E-170 | 51 | 379 | IPR010409 | GAGA-binding transcriptional activator |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 380 aa Download sequence Send to blast |
MATITFTPPM AATIVSPSLS LSHILRLKKN QAWRGDFIVA LFSWLQANVI MDNLGHRENG 60 RQRPDQYKAV HTQWMMPQRQ LKDHHSMNLL ALMNERDSAI RERDHALAEK KAAIAERDMA 120 FAQRDAAMAE RNAAVVERDN ALAALELART NGLNVNNGNE FPQGSLSGAK NIHHHDQLSH 180 VQSSPLQLAD SPYDHVREMH ILEAYPVSTA PGNIGKAKRP KKNNGQASPL KRPSGVLRKT 240 KKATSDWKNV GMSGGGEDSG RVSVMKNEWK DQDLGLNQVA FDESTMPAPA CSCTGKLRQC 300 YKWGNGGWQS SCCTMSISMY PLPVMPNKRH ARMGGRKMSG GAFTKLLSRL AAEGHDLSTP 360 VDLKDHWAKH GTNRYITIR* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional regulator that specifically binds to GA-rich elements (GAGA-repeats) present in regulatory sequences of genes involved in developmental processes. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
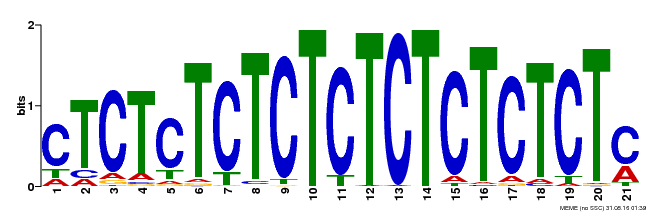
| Motif ID | Method | Source | Motif file |
| MP00540 | DAP | Transfer from AT5G42520 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | PH01000352G0060 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006655749.1 | 0.0 | PREDICTED: barley B recombinant-like protein D | ||||
| Refseq | XP_015693740.1 | 0.0 | PREDICTED: barley B recombinant-like protein D | ||||
| Swissprot | Q5VSA8 | 0.0 | BBRD_ORYSJ; Barley B recombinant-like protein D | ||||
| TrEMBL | A0A3L6DC17 | 0.0 | A0A3L6DC17_MAIZE; Barley B recombinant-like protein D | ||||
| TrEMBL | J3MAZ1 | 0.0 | J3MAZ1_ORYBR; Uncharacterized protein | ||||
| STRING | OB06G11850.1 | 0.0 | (Oryza brachyantha) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6226 | 37 | 56 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G42520.1 | 1e-102 | basic pentacysteine 6 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



