 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PH01000333G0160 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Bambusoideae; Arundinarodae; Arundinarieae; Arundinariinae; Phyllostachys
|
||||||||
| Family | HSF | ||||||||
| Protein Properties | Length: 248aa MW: 26774.6 Da PI: 8.9497 | ||||||||
| Description | HSF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HSF_DNA-bind | 102.4 | 3.9e-32 | 25 | 116 | 2 | 102 |
HHHHHHHHHCTGGGTTTSEESSSSSEEEES-HHHHHHHTHHHHSTT--HHHHHHHHHHTTEEE---SSBTTTTXTTSEEEEESXXXXXXXXXXX CS
HSF_DNA-bind 2 FlkklyeiledeelkeliswsengnsfvvldeeefakkvLpkyFkhsnfaSFvRQLnmYgFkkvkdeekkskskekiweFkhksFkkgkkelle 95
F+ k++++++d++++ +++w gn+f+vld+ f++ +Lp+yFkh+nfaSFvRQLn YgF+kv+ ++ weF+h+sF +g+ +ll
PH01000333G0160 25 FVAKTFHMVSDPATDAIVRWGGAGNTFLVLDPAAFSDFLLPSYFKHRNFASFVRQLNSYGFRKVDPDR---------WEFAHESFLRGQAQLLP 109
9****************************************************************999.........***************** PP
XXXXXXX CS
HSF_DNA-bind 96 kikrkks 102
i rkk+
PH01000333G0160 110 LIVRKKK 116
****997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.10 | 1.6E-34 | 21 | 108 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SMART | SM00415 | 5.5E-48 | 21 | 114 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| SuperFamily | SSF46785 | 1.63E-31 | 22 | 114 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| Pfam | PF00447 | 3.9E-28 | 25 | 114 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 2.8E-17 | 25 | 48 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 2.8E-17 | 63 | 75 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PROSITE pattern | PS00434 | 0 | 64 | 88 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 2.8E-17 | 76 | 88 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0006970 | Biological Process | response to osmotic stress | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 248 aa Download sequence Send to blast |
MGSECKGHEF QAAGGGGGQG AIVPFVAKTF HMVSDPATDA IVRWGGAGNT FLVLDPAAFS 60 DFLLPSYFKH RNFASFVRQL NSYGFRKVDP DRWEFAHESF LRGQAQLLPL IVRKKKAGAA 120 GRELCEEGEE VRGTIQAVQR LRDEQRGMDQ ELQTMNCRLR AAENRPGQMM AFLAKLADEP 180 GVVLRAMLAK EEELAAAGKG CAPGKKRRIG AEAGRGDRGA MAGADVAELA QGRAVPFPFS 240 VLGQVFY* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5d5u_B | 1e-26 | 7 | 114 | 11 | 129 | Heat shock factor protein 1 |
| 5d5v_B | 1e-26 | 7 | 114 | 11 | 129 | Heat shock factor protein 1 |
| 5d5v_D | 1e-26 | 7 | 114 | 11 | 129 | Heat shock factor protein 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional regulator that specifically binds DNA of heat shock promoter elements (HSE). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00379 | DAP | Transfer from AT3G24520 | Download |
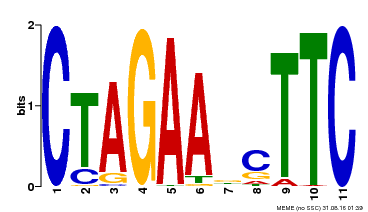
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | PH01000333G0160 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK066316 | 0.0 | AK066316.1 Oryza sativa Japonica Group cDNA clone:J013066G03, full insert sequence. | |||
| GenBank | AY344493 | 0.0 | AY344493.1 Oryza sativa (japonica cultivar-group) heat shock factor RHSF11 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015633152.1 | 1e-139 | heat stress transcription factor C-1b | ||||
| Swissprot | Q942D6 | 1e-140 | HFC1B_ORYSJ; Heat stress transcription factor C-1b | ||||
| TrEMBL | A0A0E0JNN0 | 1e-139 | A0A0E0JNN0_ORYPU; Uncharacterized protein | ||||
| STRING | OPUNC01G29770.1 | 1e-140 | (Oryza punctata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP11298 | 31 | 41 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G24520.1 | 2e-62 | heat shock transcription factor C1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



