 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PH01000210G1070 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Bambusoideae; Arundinarodae; Arundinarieae; Arundinariinae; Phyllostachys
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 370aa MW: 40807.2 Da PI: 4.4445 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 59.1 | 1e-18 | 117 | 166 | 2 | 55 |
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
+y+G+r+++ +g+W+AeIrdp++ r++lg+f +aeeAa+a++a +++++g
PH01000210G1070 117 QYRGIRQRP-WGKWAAEIRDPRK---GVRVWLGTFNSAEEAARAYDAEARRIRG 166
69*******.**********954...4************************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:3.30.730.10 | 1.1E-31 | 117 | 175 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 7.45E-32 | 117 | 176 | No hit | No description |
| SuperFamily | SSF54171 | 1.63E-21 | 117 | 174 | IPR016177 | DNA-binding domain |
| SMART | SM00380 | 7.8E-36 | 117 | 180 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 23.881 | 117 | 174 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 2.3E-11 | 118 | 166 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 4.2E-11 | 118 | 129 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 4.2E-11 | 140 | 156 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0070483 | Biological Process | detection of hypoxia | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005886 | Cellular Component | plasma membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 370 aa Download sequence Send to blast |
MCGGAILEFK EPARSRRLAE GSLWPEKKKS RRGGGERRNP VVFVEEDDED FEACFEEFEG 60 DSGDSDLEFG EEEEDDVVEI KPPAVKRAFS RDGLSTMTTA GFDGPAARSA KRTRKNQYRG 120 IRQRPWGKWA AEIRDPRKGV RVWLGTFNSA EEAARAYDAE ARRIRGKKAK VNFPEAPTVA 180 QKRHAGPIAA KAPKSSVEQK PTVKPAVNNL ANTNAFIYPS ADFTSNKLFV QPDNMPFVPA 240 MNSAAPIDDP MNIYSDQGSN SFGCSDLSWE NDTKTPDITS IAPISTIPEG DESAFIKSDT 300 HNSLVPPVME NNTVDLTDGL TDLEPYMRFL LDGGAGESID SLLNLDGSQD VVSDMDLWSF 360 DDMPISSDFY |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1gcc_A | 1e-19 | 116 | 174 | 1 | 60 | ETHYLENE RESPONSIVE ELEMENT BINDING FACTOR 1 |
| 2gcc_A | 1e-19 | 117 | 174 | 5 | 63 | ATERF1 |
| 3gcc_A | 1e-19 | 117 | 174 | 5 | 63 | ATERF1 |
| 5wx9_A | 2e-19 | 116 | 175 | 13 | 73 | Ethylene-responsive transcription factor ERF096 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcriptional activator that may be involved in defense signaling pathway. Binds in vitro to the DNA sequence 5'-AGCCGCC-3' of the GCC-box element found in pathogenesis-related (PR) gene promoters. The transcriptional activation is enhanced in vitro by the presence of MPK12/BWMK1. {ECO:0000269|PubMed:12913152}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00204 | DAP | Transfer from AT1G53910 | Download |
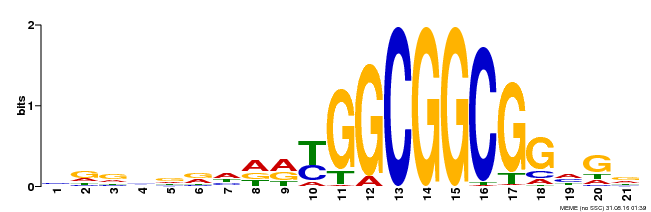
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | PH01000210G1070 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | FP100982 | 0.0 | FP100982.1 Phyllostachys edulis cDNA clone: bphylf055p09, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025816350.1 | 0.0 | ethylene-responsive transcription factor 1 | ||||
| Swissprot | Q6K7E6 | 0.0 | ERF1_ORYSJ; Ethylene-responsive transcription factor 1 | ||||
| TrEMBL | A0A3L6RNH8 | 0.0 | A0A3L6RNH8_PANMI; Ethylene-responsive transcription factor 1-like | ||||
| STRING | Pavir.Aa00281.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1039 | 38 | 134 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G53910.2 | 2e-34 | related to AP2 12 | ||||



