 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PH01000176G0870 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Bambusoideae; Arundinarodae; Arundinarieae; Arundinariinae; Phyllostachys
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 588aa MW: 64319.1 Da PI: 7.8918 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 98.8 | 3.7e-31 | 267 | 321 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
k+rlrWt eLHerF++av++L G+ekAtPk +l+lmkv+gLt++hvkSHLQkYRl
PH01000176G0870 267 KARLRWTLELHERFMDAVNKLEGPEKATPKGVLKLMKVEGLTIYHVKSHLQKYRL 321
68****************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 11.125 | 264 | 324 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 3.05E-16 | 265 | 321 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 1.1E-29 | 265 | 322 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.4E-23 | 267 | 322 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.1E-7 | 269 | 320 | IPR001005 | SANT/Myb domain |
| Pfam | PF14379 | 5.0E-22 | 357 | 403 | IPR025756 | MYB-CC type transcription factor, LHEQLE-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 588 aa Download sequence Send to blast |
MSSQSVVAVK QIAAPDKTVL GCTSTQPSVH KLFDAKLDHQ GLMDDNLSST TQSSNIKNEL 60 IRLSSLPKSL PFNVQKRSPE SDPESPPSQI SHPKFSDPIF SNSSTFCTSL FSSSSTNSEP 120 CRQMGTLPFL PHPPKCEQQV SAGQSSSSSL LLNGDIGNAH DEAEHSDDFE DFLNLSGDAS 180 DGSFHGENNA LAFAEQMEFQ LLSEQLGVAV TDNEESPRLD DIYGTPLQLS SLPVSSCSNQ 240 GLQNIGSPVK VQLSSSRSSS GSATTNKARL RWTLELHERF MDAVNKLEGP EKATPKGVLK 300 LMKVEGLTIY HVKSHLQKYR LAKYLPETKE DKKASSEDKK AHSSSSENDS GKKKNLQVAE 360 ALRMQMEVQK QLHEQLEVQR QLQLRIEEHA RYLQRILEEQ QKARSSVSLK NPTDAQAGLP 420 ESTSKEKTDS EAATTSPQPS KNRAPDADAD AECSSPVDKK RAKVHADLDF LVRASAPSLP 480 VEVVAPLGPG VPAVVLARRR RLPQRRRAPA EPAALDQDRV HLFLGAARET YGFRWGRTDL 540 GPRGFERGFR GPEAVDVVDA RWSHGGMGKI GKRGWGGCVR MISGRKW* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 2e-24 | 267 | 324 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 2e-24 | 267 | 324 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 2e-24 | 267 | 324 | 1 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 2e-24 | 267 | 324 | 1 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 2e-24 | 267 | 324 | 1 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 2e-24 | 267 | 324 | 1 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 2e-24 | 267 | 324 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 2e-24 | 267 | 324 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 2e-24 | 267 | 324 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 2e-24 | 267 | 324 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 2e-24 | 267 | 324 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 2e-24 | 267 | 324 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in phosphate starvation signaling (PubMed:26082401). Binds to P1BS, an imperfect palindromic sequence 5'-GNATATNC-3', to promote the expression of inorganic phosphate (Pi) starvation-responsive genes (PubMed:26082401). Functionally redundant with PHR1 and PHR2 in regulating Pi starvation response and Pi homeostasis (PubMed:26082401). {ECO:0000269|PubMed:26082401}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00354 | DAP | Transfer from AT3G13040 | Download |
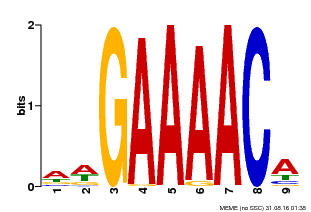
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | PH01000176G0870 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated under Pi starvation conditions. {ECO:0000269|PubMed:26082401}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025812859.1 | 0.0 | protein PHOSPHATE STARVATION RESPONSE 3 | ||||
| Refseq | XP_025812860.1 | 0.0 | protein PHOSPHATE STARVATION RESPONSE 3 | ||||
| Refseq | XP_025812862.1 | 0.0 | protein PHOSPHATE STARVATION RESPONSE 3 | ||||
| Refseq | XP_025812863.1 | 0.0 | protein PHOSPHATE STARVATION RESPONSE 3 | ||||
| Swissprot | Q6YXZ4 | 0.0 | PHR3_ORYSJ; Protein PHOSPHATE STARVATION RESPONSE 3 | ||||
| TrEMBL | A0A3L6PFL7 | 0.0 | A0A3L6PFL7_PANMI; Uncharacterized protein | ||||
| STRING | Pavir.J27649.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP4526 | 37 | 69 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G13040.2 | 6e-70 | G2-like family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



