 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PH01000143G0710 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Bambusoideae; Arundinarodae; Arundinarieae; Arundinariinae; Phyllostachys
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 1102aa MW: 122366 Da PI: 7.7902 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 181.1 | 2.8e-56 | 458 | 585 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLp.k.kvkaeekewyfFskrdkkyatgkrknratksgyWkatgkdkev 92
lppGfrFhPtdeel+ +yLk+k++g+++el e+i+evd+yk+ePwdLp k + +++ ewyfFs+rd+ky++g+r+nratk+gyWkatgkd++v
PH01000143G0710 458 LPPGFRFHPTDEELIIYYLKRKINGRQIEL-EIIPEVDLYKCEPWDLPeKsFLPSKDLEWYFFSPRDRKYPNGSRTNRATKAGYWKATGKDRKV 550
79****************************.99**************95334455777************************************ PP
NAM 93 lskkgelvglkktLvfykgrapkgektdWvmheyrl 128
s +++ vg+kktLv+y+grap+g++tdWvmheyrl
PH01000143G0710 551 NS-QRRPVGMKKTLVYYRGRAPHGSRTDWVMHEYRL 585
**.999****************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF57756 | 1.13E-7 | 97 | 120 | IPR001878 | Zinc finger, CCHC-type |
| SMART | SM00343 | 5.8E-5 | 100 | 116 | IPR001878 | Zinc finger, CCHC-type |
| PROSITE profile | PS50158 | 11.004 | 101 | 115 | IPR001878 | Zinc finger, CCHC-type |
| Gene3D | G3DSA:4.10.60.10 | 3.7E-7 | 101 | 126 | IPR001878 | Zinc finger, CCHC-type |
| Pfam | PF00098 | 6.7E-7 | 101 | 116 | IPR001878 | Zinc finger, CCHC-type |
| Pfam | PF13976 | 1.3E-7 | 332 | 380 | IPR025724 | GAG-pre-integrase domain |
| SuperFamily | SSF101941 | 2.48E-64 | 451 | 608 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 61.367 | 458 | 608 | IPR003441 | NAC domain |
| Pfam | PF02365 | 4.0E-29 | 459 | 585 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1102 aa Download sequence Send to blast |
MGENQGLQQV LEKLAQLLTE KTDCISTVKK WIERRRVIEF LKGLNPRFDG RRATMFHQPT 60 LPTLEEAIAA IAQEKTRLKV MTSTASQITR PVFAVTKTRD CFNCGEMGHL SRNCPHPRRP 120 NRGKDRVYVK SGQKGSGSKG GKSSYKANFA MTGEESPERV EVSVAELEEL RQLKKRVENL 180 GNKDKGTAIV SSASANLAHS NPGKWIIDSG ASTHVTGMLD EFKSYTPFSP TYMETIQTAD 240 GTVQPIKGVG IVQCSPSITL SSVLHVPSFL VNLLSISSLV DHLDCWVIFD RETCLIQERQ 300 NGRQLGTGTR SNGLWYLDPR EGDVGLCTAL VATSSEEEAR VMLMHCRLDH ISFETMSKVF 360 PDVINKVDRS KLMCDTCEFA HLQSAHPSGI IVPTGSSVTQ LLTVGMQPSQ QSVRTVIHSR 420 TPEQTALDSL ACLPSSKHPL AVRVLRGRSR KQMAPVSLPP GFRFHPTDEE LIIYYLKRKI 480 NGRQIELEII PEVDLYKCEP WDLPEKSFLP SKDLEWYFFS PRDRKYPNGS RTNRATKAGY 540 WKATGKDRKV NSQRRPVGMK KTLVYYRGRA PHGSRTDWVM HEYRLDEREC ETDTGLQDAY 600 ALCRVFKKTA PGPKIIEHYG AVHHHLEQPQ WMASSVDRSP TLHVSCDGRG DDFESSSFSF 660 PTEVPMDSMH GGFGMQMGSA PHEDGKWMQF LSEDAFNATN PFFMNPAASN FSCLPSKVDV 720 ALECARLQHR LSLPPLEVED FPQDVSLDTK TGILRSNPNE VDILQEFLSV ASASQELING 780 SSSGYPEMWL GAGTSSASTH YMNELSSLVE LGVKTKEEVD SFYQMGGIGT SGHDEPARLV 840 EITDMEFKEE KKKVENLRGV KLVNNDLGEI VVGGDGSIPT EGITHYPVKD TTDNSGGTDT 900 APIFSQSQPD DFAIGFNDVN PNASFDLYEK VDVKHGLFVS RVGAAKTFFH RTEPSKKVSF 960 HLNPVASDVS EAIEKFHFPI TTKISGRVSI FSKFKALIMD KFLMRPSYKR YLGSKETTAS 1020 ELLQIMSLLL APKEVTGPTE QELVKKKAKE VMKPGPGWGC DGGNNVWLPL SKGRGISSMF 1080 LSGKWTFLTP ALAIRTPGCN H* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 3ulx_A | 3e-55 | 448 | 608 | 5 | 168 | Stress-induced transcription factor NAC1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00212 | DAP | Transfer from AT1G65910 | Download |
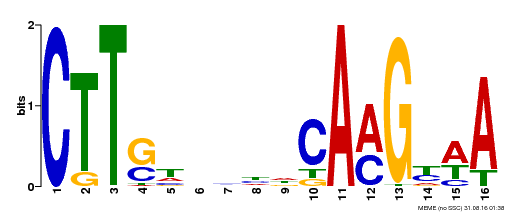
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | PH01000143G0710 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB265821 | 0.0 | AB265821.1 Oryza sativa Japonica Group RIM1 mRNA for NAC protein, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006649289.1 | 0.0 | PREDICTED: NAC domain-containing protein 86-like | ||||
| TrEMBL | A0A0E0K812 | 0.0 | A0A0E0K812_ORYPU; Uncharacterized protein | ||||
| STRING | OPUNC03G01460.1 | 0.0 | (Oryza punctata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6624 | 36 | 51 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G65910.1 | 1e-127 | NAC domain containing protein 28 | ||||



