 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PH01000029G0970 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Bambusoideae; Arundinarodae; Arundinarieae; Arundinariinae; Phyllostachys
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 466aa MW: 51118.3 Da PI: 7.5037 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 30.8 | 6.5e-10 | 179 | 221 | 3 | 45 |
XXCHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 3 elkrerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaL 45
+ k rr+++NReAAr+sR+RKka+i++Le +L++ ++L
PH01000029G0970 179 DHKTLRRLAQNREAARKSRLRKKAYIQNLESSRLKLTQLEQEL 221
56889*************************9777777666555 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.20.5.170 | 1.6E-8 | 170 | 224 | No hit | No description |
| SMART | SM00338 | 1.3E-9 | 177 | 240 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 9.197 | 179 | 223 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF00170 | 4.5E-7 | 180 | 221 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 2.81E-7 | 181 | 224 | No hit | No description |
| CDD | cd14708 | 2.33E-22 | 182 | 233 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 184 | 199 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF14144 | 1.6E-33 | 263 | 338 | IPR025422 | Transcription factor TGA like domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009410 | Biological Process | response to xenobiotic stimulus | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005737 | Cellular Component | cytoplasm | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 466 aa Download sequence Send to blast |
MESRRAGGAA AAAEDVRGKV PGFGTPQHTI HTDVNSMQPS RVTDFGALAH SAGFRIEDLA 60 NFSTNTLFNL KPNTHTIIDD HLQFGNHGKS ISPTDITTTA AAAVDPQALV QQKGAQPNLV 120 ALRTGNIENW GESTMAATSP RTDTSTDPDI DERNQLFEQG QLAAPTASDS SDKSKDKSDH 180 KTLRRLAQNR EAARKSRLRK KAYIQNLESS RLKLTQLEQE LQRARQQGVF ISTSGDQPHS 240 MSGNGALAFD MEYARWLEEH NKHINELRAA VNAHAGDNDL RSIVDSIMAH YDEIFRLKGV 300 AAKADVFHVL SGMWKTPAER CFMWLGGFRS SELLKLLAGQ LEPLTEQQLT GICNLQQSSQ 360 QAEDALSQGM EALQQSLAET LASGSLGPAG SSGNVANYMG HMAMAMGKLG TLENFLRQAD 420 NLRLQTLQQM QRILTTRQSA RALLAISDYF SRLRALSSLW LARPRE |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-TGACG-3'. Recognizes ocs elements like the as-1 motif of the cauliflower mosaic virus 35S promoter. Binding to the as-1-like cis elements mediate auxin- and salicylic acid-inducible transcription. Binds to the hexamer motif 5'-ACGTCA-3' of histone gene promoters. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00492 | DAP | Transfer from AT5G06950 | Download |
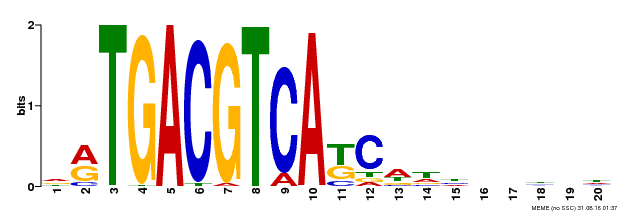
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | PH01000029G0970 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | FP097013 | 0.0 | FP097013.1 Phyllostachys edulis cDNA clone: bphyem208m15, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002458658.1 | 0.0 | transcription factor TGAL1 | ||||
| Swissprot | Q41558 | 0.0 | HBP1C_WHEAT; Transcription factor HBP-1b(c1) (Fragment) | ||||
| TrEMBL | F2DGH6 | 0.0 | F2DGH6_HORVV; Predicted protein | ||||
| STRING | Sb03g037560.1 | 0.0 | (Sorghum bicolor) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP486 | 38 | 185 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G06950.4 | 1e-140 | bZIP family protein | ||||



