 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PH01000000G0910 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Bambusoideae; Arundinarodae; Arundinarieae; Arundinariinae; Phyllostachys
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 534aa MW: 58971.2 Da PI: 7.3043 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 51.1 | 3e-16 | 59 | 103 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT++E++l+++a +++G W++I +++g ++t+ q++s+ qk+
PH01000000G0910 59 REKWTEDEHKLFLEALQLHGRA-WRRIQEHIG-TKTAVQIRSHAQKF 103
789*****************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 3.59E-15 | 53 | 106 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 20.318 | 54 | 108 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 2.6E-15 | 57 | 106 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 3.8E-13 | 58 | 106 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.3E-14 | 59 | 102 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.6E-9 | 59 | 101 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 4.55E-10 | 61 | 104 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 534 aa Download sequence Send to blast |
MAPFQEIEAS DDQRPVADHV GHQNLMENLG NPLNSSDMDM LAEARVPKAR KPYTITKQRE 60 KWTEDEHKLF LEALQLHGRA WRRIQEHIGT KTAVQIRSHA QKFFSKVIRE SSGDNSNSSG 120 VAPPIQIPPP RPKRKPVHPY PRKLGNAPGK LVPVLRQLEK PQLQIQTLCE QEKGSPTSVL 180 TTTQKGYETL GSDSGESPAS TIDNEERCPT SSVATAELAV QVPPTDVKEA KGNTTSKEAV 240 RNRSEASVLR LFGKRVVVND LHQKANRSAG NLQNAADMEL DASVETPTSG TGKFASHGAA 300 EANTWTPWLI DTRQFLYYLP QGEVFSMHSA LPCFSYNNGS VPCPPLLNPK AVAQNQQHQH 360 QPSQAVNYKF MQREGSWTES NTASSSVPET ATQNSDSAES CRKANGSEDE MVPVPGLRKC 420 VSPASICQRG FVPYKRCAAE SEVLQSQMTR TDVLREGRSA VTDERQIETT LVFHNWKSSC 480 GPCLPMYSSN LWYFPFSSLP VCNFAPMLEV QTSEVNGLSC SDLYFERVVV GLY* |
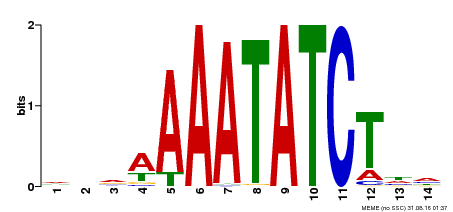
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00515 | DAP | Transfer from AT5G17300 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | PH01000000G0910 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | FO203437 | 0.0 | FO203437.1 Phyllostachys heterocycla genomic DNA, BAC clone: PH01B001I13, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015636284.1 | 0.0 | uncharacterized protein LOC4336779 | ||||
| TrEMBL | L0P1M2 | 0.0 | L0P1M2_9POAL; PH01B001I13.25 protein | ||||
| STRING | ONIVA04G22180.1 | 0.0 | (Oryza nivara) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2722 | 36 | 86 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G37260.1 | 3e-33 | MYB_related family protein | ||||



