 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pahal.I04064.2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 508aa MW: 52469.8 Da PI: 6.5812 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 50.3 | 4.3e-16 | 304 | 349 | 5 | 55 |
HHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 5 hnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
hne Er+RRdriN+++ +L++l+P++ K +Ka++L ++++Y+k+Lq
Pahal.I04064.2 304 HNESERKRRDRINQKMKTLQKLVPNS-----SKTDKASMLDEVIDYLKQLQ 349
*************************8.....7******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.280.10 | 4.6E-18 | 297 | 365 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 18.025 | 299 | 348 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 1.16E-19 | 302 | 372 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 1.2E-13 | 304 | 349 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 6.00E-18 | 304 | 353 | No hit | No description |
| SMART | SM00353 | 1.2E-18 | 305 | 354 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009567 | Biological Process | double fertilization forming a zygote and endosperm | ||||
| GO:0009506 | Cellular Component | plasmodesma | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 508 aa Download sequence Send to blast |
MNQCVPSWDL DDPALVAAGG GGLNNHPVPA AGGGHHRGGA APAGTTATAG GAFAPVVVPM 60 ADQYYEVAEL TWEKGNISSH GLLNRPPPSK YHHQPAAPPP SHLQGIGGGG GAAGDRETLE 120 AVVGEAAARS HFLSQPPVHL AHWIEVGVGA AAAAVARPGA DALVPCAARA EEAAEGADAA 180 DASRRKRARV VGEDGLVCAS QGSAAPGRRG DSALITLDGC GTGADDVCGF TTTTNNSTSL 240 EREDKGSPDT ENTSIGGGAS DSRCFSRRSQ RDGLCDEGEN VVTTGDGAVR SSVSTKRSRA 300 AAIHNESERK RRDRINQKMK TLQKLVPNSS KTDKASMLDE VIDYLKQLQA QVQVMSRMST 360 MMMPMAMPQL QMSAVMAQMA QMAQMAQMAQ GMMNMGSLAQ PAAAYAGLTP PMMPTFVPTM 420 PWDPTTSGAG SVVGTGTADR APQPPAAVAG AVPPDAFSAF LACQAQQQSG QQQQQQPGSM 480 EAYNKMVALY QKMSQQQQGQ PSSSSKQ* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 307 | 312 | ERKRRD |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00034 | PBM | Transfer from AT4G00050 | Download |
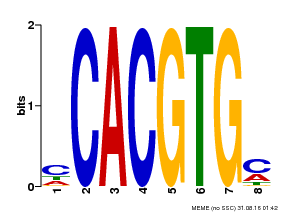
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025792489.1 | 0.0 | transcription factor UNE10 isoform X3 | ||||
| TrEMBL | A0A2S3INT7 | 0.0 | A0A2S3INT7_9POAL; Uncharacterized protein | ||||
| STRING | Pavir.Ia02188.1.p | 0.0 | (Panicum virgatum) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00050.1 | 7e-31 | bHLH family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pahal.I04064.2 |



