 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pahal.I00784.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1032aa MW: 115109 Da PI: 5.7907 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 178.2 | 1e-55 | 19 | 136 | 2 | 118 |
CG-1 2 lke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfq 95
lke ++rwl++ ei++iL+n+++++++ e+++rp+sgsl+L++rk++ryfrkDG++w+kk+dgktv+E+he+LK g+++vl+cyYah+een++fq
Pahal.I00784.1 19 LKEaRHRWLRPAEICEILKNYRNFRIAPEPPNRPPSGSLFLFDRKVLRYFRKDGHNWRKKRDGKTVKEAHERLKSGSIDVLHCYYAHGEENENFQ 113
45559****************************************************************************************** PP
CG-1 96 rrcywlLeeelekivlvhylevk 118
rr+yw+Lee++ +ivlvhylevk
Pahal.I00784.1 114 RRSYWMLEEDFMHIVLVHYLEVK 136
********************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 83.085 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 7.9E-78 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 2.4E-49 | 21 | 135 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 1.7E-7 | 442 | 527 | IPR013783 | Immunoglobulin-like fold |
| CDD | cd00102 | 2.64E-4 | 442 | 529 | No hit | No description |
| Pfam | PF01833 | 3.3E-10 | 442 | 522 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 1.38E-18 | 443 | 528 | IPR014756 | Immunoglobulin E-set |
| CDD | cd00204 | 1.63E-12 | 623 | 735 | No hit | No description |
| SuperFamily | SSF48403 | 5.29E-16 | 634 | 737 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 1.9E-16 | 637 | 738 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 16.998 | 643 | 737 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 8.576 | 643 | 675 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 8.736 | 676 | 708 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 2.92E-8 | 848 | 900 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.07 | 849 | 871 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.169 | 850 | 879 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0029 | 851 | 870 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0022 | 872 | 894 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.578 | 873 | 897 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.0E-4 | 875 | 894 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1032 aa Download sequence Send to blast |
MAEARRYAIA PQLDVDQILK EARHRWLRPA EICEILKNYR NFRIAPEPPN RPPSGSLFLF 60 DRKVLRYFRK DGHNWRKKRD GKTVKEAHER LKSGSIDVLH CYYAHGEENE NFQRRSYWML 120 EEDFMHIVLV HYLEVKGGKS SSRIRGHDDM LQAARTDSPL SQLPSQTTEG ESSVSGQATE 180 YEETESDIYS GGAGYHPFSW TQQHENGGGP VMGASILSCY IPSPFVGNHQ GFLSTATTTD 240 LYSHGQDALP VALNEPALGI ALNGADNQLD PSSLNGLPDQ GIHRMPPPQI TDPSKQFPFT 300 EGPGIESFTF GEVYNGLGIK DADTVDTDEE SLWQLPGAIS SFPTEDSFQQ NGRSLEETIN 360 YPLLKTQSSG LSDILKDSFK KSDSFTRWMS KELGEVDDSQ IRSSSGVYWN SEETDNIIEA 420 SSRDQLDQFT VDPVLAQDQL FSISDFSPSW TYAGSKTRVL ITGRFLNSYE VQRCKWSCMF 480 GEVEVPAEIS ADGTLRCYSP SHKPGRVPFY VTCSNRLACS EIREFEFRPS NSKHMDVPSP 540 HDDANKTYLQ MRLDDLLSLG QDEYQATVSN PTKEMIDLSK KISSLMTDND SWSELLKLAC 600 DNELATDDKQ DQFFENRLKE KLHIWLVHKA GDGGKGPSVL DEEGQGVLHL AAALGYDWAI 660 RPTISAGVSI NFRDAHGWTA LHWAAFCGRE RTVVALIALG AAPGALTDPT LDFPTGSTPA 720 DLASANGYKG ISGFLAESAL ISHLQTLDLK ETMGSNASEI SGLPGIGDVT ERRASPLAGE 780 GLLAGSMGDS LGAVRNAAQA AARIYQVFRM QSFQRKQAVQ YEDDNGAISD DRALSLLSVK 840 PSKPGQLDPL HAAATRIQNK YRGWKGRKEF LLIRQRIVKI QAHVRGHQVR KHYRKIIWSV 900 GIVEKIILRW RRKGAGLRGF RSTEGAVEGT SSSRSDLIQN KPAQDDYDFL QQGRKQTEER 960 LQKALARVKS MVQYPDARDQ YQRILTVVTR LQESQALQEK MLESSTDMDE GFVMSEFQKL 1020 WDDDMPMPGN I* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
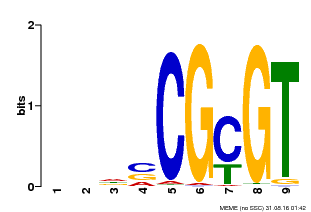
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025793890.1 | 0.0 | calmodulin-binding transcription activator 3-like isoform X2 | ||||
| TrEMBL | A0A2S3ITE5 | 0.0 | A0A2S3ITE5_9POAL; Uncharacterized protein | ||||
| STRING | Si034076m | 0.0 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP608 | 38 | 140 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pahal.I00784.1 |



