 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pahal.A02949.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 457aa MW: 48450.9 Da PI: 8.5964 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 50.6 | 4.4e-16 | 51 | 95 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT++E+ ++++a +++G W++I +++g ++t+ q++s+ qk+
Pahal.A02949.1 51 REKWTEDEHRRFLEALQLHGRA-WRRIQEHIG-TKTAVQIRSHAQKF 95
789*****************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.69E-15 | 45 | 99 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 19.732 | 46 | 100 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 3.4E-16 | 49 | 98 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 1.7E-12 | 50 | 98 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.8E-9 | 51 | 91 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 6.5E-14 | 51 | 94 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 8.70E-10 | 53 | 96 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 457 aa Download sequence Send to blast |
MACLEEHQMA TDEVAVAVDH RKDPLDSGDM DLSGEEHVPK ARKPYTITKQ REKWTEDEHR 60 RFLEALQLHG RAWRRIQEHI GTKTAVQIRS HAQKFFTKVV RESSGSSTTA GAAPAIQIPP 120 PRPKRKPAHP YPRKVDGAAK KHVPALRQLE KPPLRTHSPR DQDDGSPTSV LTTARTVLRA 180 EELGSVFANS SSGSRTPSQS AAGSDEHGNG GGSPASTINR EDGGVSPSVA TDELATRTAN 240 AKVFGDAKEA GTEAPVFKLF GKKVVVKDSY GHPESGRDLK IGASPAIVAA GGSSWNPWPG 300 AVQQLMCFVP QPDGFAAQSV VPWLAYNGSL PCALFYPQAA PSAQQLHQTS EPPDHKRTQR 360 EGSRTGSNTA SSAVPAAQNS DATESHGPGQ ETTSESGAVP PAAAAVPVPR LARCASSASF 420 SRRGFVPHKR CAAESEAPRP VAAGEEADGE LTRLCL* |
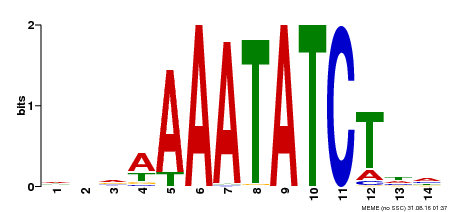
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00515 | DAP | Transfer from AT5G17300 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025814514.1 | 0.0 | protein REVEILLE 1-like | ||||
| TrEMBL | A0A2S3GS63 | 0.0 | A0A2S3GS63_9POAL; Uncharacterized protein | ||||
| STRING | Pavir.Aa00960.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2722 | 36 | 86 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G01060.3 | 8e-35 | MYB_related family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pahal.A02949.1 |



