 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pahal.A02530.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | AP2 | ||||||||
| Protein Properties | Length: 684aa MW: 71992.5 Da PI: 6.2946 | ||||||||
| Description | AP2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 44.7 | 3.4e-14 | 280 | 339 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrd.pse.ng..kr.krfslgkfgtaeeAakaaiaarkkleg 55
s y+GV++++++gr++A+++d + g ++ ++++lg ++ +++Aa+a++ a++k++g
Pahal.A02530.1 280 SIYRGVTRHRWTGRYEAHLWDnSCRrEGqsRKgRQVYLGGYDKEDKAARAYDLAALKYWG 339
57*******************666664477446**********99*************98 PP
| |||||||
| 2 | AP2 | 49.8 | 8.7e-16 | 382 | 433 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s+y+GV+++++ grW A+I + +k +lg+f t+eeAa+a++ a+ k++g
Pahal.A02530.1 382 SKYRGVTRHHQHGRWQARIGRVAG---NKDLYLGTFSTEEEAAEAYDIAAIKFRG 433
89****************988532...5************************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 1.18E-16 | 280 | 349 | IPR016177 | DNA-binding domain |
| CDD | cd00018 | 3.21E-20 | 280 | 349 | No hit | No description |
| Pfam | PF00847 | 2.9E-11 | 280 | 339 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 8.1E-15 | 281 | 348 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 18.861 | 281 | 347 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 1.0E-28 | 281 | 353 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.6E-6 | 282 | 293 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 5.56E-18 | 382 | 443 | IPR016177 | DNA-binding domain |
| CDD | cd00018 | 9.91E-24 | 382 | 443 | No hit | No description |
| Pfam | PF00847 | 3.1E-10 | 382 | 433 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 2.5E-18 | 383 | 442 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.467 | 383 | 441 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 8.1E-34 | 383 | 447 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.6E-6 | 423 | 443 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000723 | Biological Process | telomere maintenance | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007389 | Biological Process | pattern specification process | ||||
| GO:0010073 | Biological Process | meristem maintenance | ||||
| GO:0010449 | Biological Process | root meristem growth | ||||
| GO:0019827 | Biological Process | stem cell population maintenance | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 684 aa Download sequence Send to blast |
MASANNWLGF SLSGQDNPQP NQDSSPAAGI DISGASDFYG LPTQQGSDGH LGVPGLRDDH 60 ASYGIMEAFN RGQQETQDWN MRGLDYNGGA SELSMLVGSS GGAGGGTGKR AVEDSEPKLE 120 DFLGGNSFVS EQDQSGGYLF SGVPMASSTN SNSGSNTMEL SMIKTWLRNN QVPQPQPPAT 180 HQPQPEEMST DASASSFGCS DSLGRNGAVA GGSSQSLALS MSTGSHLPMV VAGGGNASGA 240 AASDSTSSEN KRASGAMDSP SSAIEAVPRK SIDTFGQRTS IYRGVTRHRW TGRYEAHLWD 300 NSCRREGQSR KGRQVYLGGY DKEDKAARAY DLAALKYWGT TTTTNFPISN YEKELEEMKH 360 MTRQEYIAYL RRNSSGFSRG ASKYRGVTRH HQHGRWQARI GRVAGNKDLY LGTFSTEEEA 420 AEAYDIAAIK FRGLNAVTNF DMSRYDVKSI LESSTLPVGG AARRLKDAVD HVEAAGATIW 480 RADMEGAGVI SQLADAGMGA YASYGHHGWP TIAFQQPSPL TVHYPYGQAP SRGWCKPEQD 540 AAVAAAAHSL QDLQQLHLGS AAHNFFQASS SSTVYNGGGG ASAAGYHQGL GGGGGSFIMP 600 SSTVVADQGH SSTANQGSTC SYGDDQEGKL IGYDAMVATA AGGDPYAAAR SGYQFSQGSG 660 STVSIARANG YSNNWSSPFN GMG* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that promotes cell proliferation, differentiation and morphogenesis, especially during embryogenesis. {ECO:0000269|PubMed:12172019}. | |||||
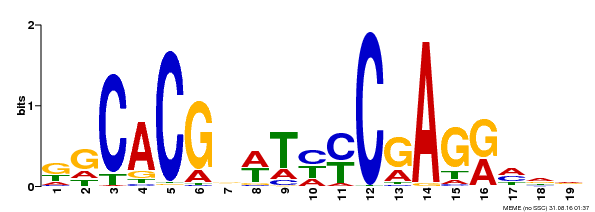
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00368 | DAP | Transfer from AT3G20840 | Download |
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | EU955732 | 0.0 | EU955732.1 Zea mays clone 1543867 protein BABY BOOM 1 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025824559.1 | 0.0 | AP2-like ethylene-responsive transcription factor BBM1 isoform X2 | ||||
| Swissprot | Q8LSN2 | 1e-136 | BBM2_BRANA; AP2-like ethylene-responsive transcription factor BBM2 | ||||
| TrEMBL | A0A2S3GQY0 | 0.0 | A0A2S3GQY0_9POAL; Uncharacterized protein | ||||
| STRING | Pavir.J07914.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1569 | 37 | 110 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G20840.1 | 1e-125 | AP2 family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pahal.A02530.1 |



